Temporal coordination of gene networks by Zelda in the early Drosophila embryo
- PMID: 22028675
- PMCID: PMC3197689
- DOI: 10.1371/journal.pgen.1002339
Temporal coordination of gene networks by Zelda in the early Drosophila embryo
Abstract
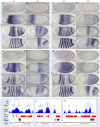
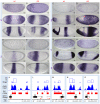
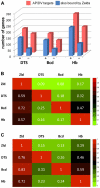
In past years, much attention has focused on the gene networks that regulate early developmental processes, but less attention has been paid to how multiple networks and processes are temporally coordinated. Recently the discovery of the transcriptional activator Zelda (Zld), which binds to CAGGTAG and related sequences present in the enhancers of many early-activated genes in Drosophila, hinted at a mechanism for how batteries of genes could be simultaneously activated. Here we use genome-wide binding and expression assays to identify Zld target genes in the early embryo with the goal of unraveling the gene circuitry regulated by Zld. We found that Zld binds to genes involved in early developmental processes such as cellularization, sex determination, neurogenesis, and pattern formation. In the absence of Zld, many target genes failed to be activated, while others, particularly the patterning genes, exhibited delayed transcriptional activation, some of which also showed weak and/or sporadic expression. These effects disrupted the normal sequence of patterning-gene interactions and resulted in highly altered spatial expression patterns, demonstrating the significance of a timing mechanism in early development. In addition, we observed prevalent overlap between Zld-bound regions and genomic "hotspot" regions, which are bound by many developmental transcription factors, especially the patterning factors. This, along with the finding that the most over-represented motif in hotspots, CAGGTA, is the Zld binding site, implicates Zld in promoting hotspot formation. We propose that Zld promotes timely and robust transcriptional activation of early-gene networks so that developmental events are coordinated and cell fates are established properly in the cellular blastoderm embryo.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
References
-
- ten Bosch JR, Benavides JA, Cline TW. The TAGteam DNA motif controls the timing of Drosophila pre-blastoderm transcription. Development. 2006;133:1967–1977. - PubMed
-
- De Renzis SD, Elemento O, Tavazoie S, Wieschaus EF. Unmasking activation of the zygotic genome using chromosomal deletions in the Drosophila embryo. PLoS Biol. 2007;5:e117. doi: 10.1371/journal.pbio.0050117. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

