Signal peptidase I: cleaving the way to mature proteins
- PMID: 22031009
- PMCID: PMC3323777
- DOI: 10.1002/pro.757
Signal peptidase I: cleaving the way to mature proteins
Abstract
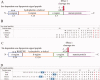
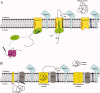
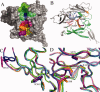
Signal peptidase I (SPase I) is critical for the release of translocated preproteins from the membrane as they are transported from a cytoplasmic site of synthesis to extracytoplasmic locations. These proteins are synthesized with an amino-terminal extension, the signal sequence, which directs the preprotein to the Sec- or Tat-translocation pathway. Recent evidence indicates that the SPase I cleaves preproteins as they emerge from either pathway, though the steps involved are unclear. Now that the structure of many translocation pathway components has been elucidated, it is critical to determine how these components work in concert to support protein translocation and cleavage. Molecular modeling and NMR studies have provided insight on how the preprotein docks on SPase I in preparation for cleavage. This is a key area for future work since SPase I enzymes in a variety of species have now been identified and the inhibition of these enzymes by antibiotics is being pursued. The eubacterial SPase I is essential for cell viability and belongs to a unique group of serine endoproteases which utilize a Ser-Lys catalytic dyad instead of the prototypical Ser-His-Asp triad used by eukaryotes. As such, SPase I is a desirable antimicrobial target. Advances in our understanding of how the preprotein interfaces with SPase I during the final stages of translocation will facilitate future development of inhibitors that display a high efficacy against SPase I function.
Copyright © 2011 The Protein Society.
Figures

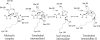
References
-
- Dalbey RE, Wickner W. Leader peptidase catalyzes the release of exported proteins from the outer surface of the Escherichia coli plasma membrane. J Biol Chem. 1985;260:15925–15931. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

