Rapid, reliable, and reproducible molecular sub-grouping of clinical medulloblastoma samples
- PMID: 22057785
- PMCID: PMC3306784
- DOI: 10.1007/s00401-011-0899-7
Rapid, reliable, and reproducible molecular sub-grouping of clinical medulloblastoma samples
Abstract
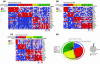
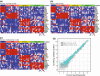
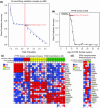
The diagnosis of medulloblastoma likely encompasses several distinct entities, with recent evidence for the existence of at least four unique molecular subgroups that exhibit distinct genetic, transcriptional, demographic, and clinical features. Assignment of molecular subgroup through routine profiling of high-quality RNA on expression microarrays is likely impractical in the clinical setting. The planning and execution of medulloblastoma clinical trials that stratify by subgroup, or which are targeted to a specific subgroup requires technologies that can be economically, rapidly, reliably, and reproducibly applied to formalin-fixed paraffin embedded (FFPE) specimens. In the current study, we have developed an assay that accurately measures the expression level of 22 medulloblastoma subgroup-specific signature genes (CodeSet) using nanoString nCounter Technology. Comparison of the nanoString assay with Affymetrix expression array data on a training series of 101 medulloblastomas of known subgroup demonstrated a high concordance (Pearson correlation r = 0.86). The assay was validated on a second set of 130 non-overlapping medulloblastomas of known subgroup, correctly assigning 98% (127/130) of tumors to the appropriate subgroup. Reproducibility was demonstrated by repeating the assay in three independent laboratories in Canada, the United States, and Switzerland. Finally, the nanoString assay could confidently predict subgroup in 88% of recent FFPE cases, of which 100% had accurate subgroup assignment. We present an assay based on nanoString technology that is capable of rapidly, reliably, and reproducibly assigning clinical FFPE medulloblastoma samples to their molecular subgroup, and which is highly suited for future medulloblastoma clinical trials.
Figures

References
-
- Cho YJ, Tsherniak A, Tamayo P, Santagata S, Ligon A, Greulich H, Berhoukim R, Amani V, Goumnerova L, Eberhart CG, Lau CC, Olson JM, Gilbertson RJ, Gajjar A, Delattre O, Kool M, Ligon K, Meyerson M, Mesirov JP, Pomeroy SL. Integrative genomic analysis of medulloblastoma identifies a molecular subgroup that drives poor clinical outcome. J Clin Oncol. 2011;29:1424–1430. doi: 10.1200/JCO.2010.28.5148. - DOI - PMC - PubMed
-
- Clifford SC, Lusher ME, Lindsey JC, Langdon JA, Gilbertson RJ, Straughton D, Ellison DW. Wnt/Wingless pathway activation and chromosome 6 loss characterize a distinct molecular sub-group of medulloblastomas associated with a favorable prognosis. Cell Cycle. 2006;5:2666–2670. doi: 10.4161/cc.5.22.3446. - DOI - PubMed
-
- Ellison DW, Dalton J, Kocak M, Nicholson SL, Fraga C, Neale G, Kenney AM, Brat DJ, Perry A, Yong WH, Taylor RE, Bailey S, Clifford SC, Gilbertson RJ. Medulloblastoma: clinicopathological correlates of SHH, WNT and non-SHH/WNT molecular subgroups. Acta Neuropathol. 2011;121:381–396. doi: 10.1007/s00401-011-0800-8. - DOI - PMC - PubMed
-
- Gajjar A, Chintagumpala M, Ashley D, Kellie S, Kun LE, Merchant TE, Woo S, Wheeler G, Ahern V, Krasin MJ, Fouladi M, Broniscer A, Krance R, Hale GA, Stewart CF, Dauser R, Sanford RA, Fuller C, Lau C, Boyett JM, Wallace D, Gilbertson RJ. Risk-adapted craniospinal radiotherapy followed by high-dose chemotherapy and stem-cell rescue in children with newly diagnosed medulloblastoma (St Jude Medulloblastoma-96): long-term results from a prospective, multicentre trial. Lancet Oncol. 2006;7:813–820. doi: 10.1016/S1470-2045(06)70867-1. - DOI - PubMed
-
- Geiss GK, Bumgarner RE, Birditt B, Dahl T, Dowidar N, Dunaway DL, Fell HP, Ferree S, George RD, Grogan T, James JJ, Maysuria M, Mitton JD, Oliveri P, Osborn JL, Peng T, Ratcliffe AL, Webster PJ, Davidson EH, Hood L, Dimitrov K. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat Biotechnol. 2008;26:317–325. doi: 10.1038/nbt1385. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

