An alternative path for the evolution of biological nitrogen fixation
- PMID: 22065963
- PMCID: PMC3207485
- DOI: 10.3389/fmicb.2011.00205
An alternative path for the evolution of biological nitrogen fixation
Abstract
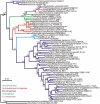
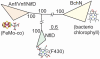
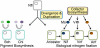
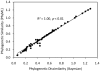
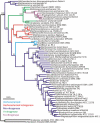
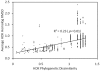
Nitrogenase catalyzed nitrogen fixation is the process by which life converts dinitrogen gas into fixed nitrogen in the form of bioavailable ammonia. The most common form of nitrogenase today requires a complex metal cluster containing molybdenum (Mo), although alternative forms exist which contain vanadium (V) or only iron (Fe). It has been suggested that Mo-independent forms of nitrogenase (V and Fe) were responsible for N(2) fixation on early Earth because oceans were Mo-depleted and Fe-rich. Phylogenetic- and structure-based examinations of multiple nitrogenase proteins suggest that such an evolutionary path is unlikely. Rather, our results indicate an evolutionary path whereby Mo-dependent nitrogenase emerged within the methanogenic archaea and then gave rise to the alternative forms suggesting that they arose later, perhaps in response to local Mo limitation. Structural inferences of nitrogenase proteins and related paralogs suggest that the ancestor of all nitrogenases had an open cavity capable of binding metal clusters which conferred reactivity. The evolution of the nitrogenase ancestor and its associated bound metal cluster was controlled by the availability of fixed nitrogen in combination with local environmental factors that influenced metal availability until a point in Earth's geologic history where the most desirable metal, Mo, became sufficiently bioavailable to bring about and refine the solution (Mo-nitrogenase) we see perpetuated in extant biology.
Keywords: evolution; great oxidation event; iron; metalloenzyme; nitrogenase; oxygen molybdenum; sulfide; vanadium.
Figures
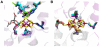
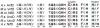
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

