Interplay between single resistance-associated mutations in the HIV-1 protease and viral infectivity, protease activity, and inhibitor sensitivity
- PMID: 22083488
- PMCID: PMC3264268
- DOI: 10.1128/AAC.05549-11
Interplay between single resistance-associated mutations in the HIV-1 protease and viral infectivity, protease activity, and inhibitor sensitivity
Abstract
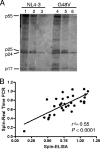
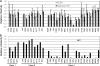
Resistance-associated mutations in the HIV-1 protease modify viral fitness through changes in the catalytic activity and altered binding affinity for substrates and inhibitors. In this report, we examine the effects of 31 mutations at 26 amino acid positions in protease to determine their impact on infectivity and protease inhibitor sensitivity. We found that primary resistance mutations individually decrease fitness and generally increase sensitivity to protease inhibitors, indicating that reduced virion-associated protease activity reduces virion infectivity and the reduced level of per virion protease activity is then more easily titrated by a protease inhibitor. Conversely, mutations at more variable positions (compensatory mutations) confer low-level decreases in sensitivity to all protease inhibitors with little effect on infectivity. We found significant differences in the observed effect on infectivity with a pseudotype virus assay that requires the protease to cleave the cytoplasmic tail of the amphotropic murine leukemia virus (MuLV) Env protein. Additionally, we were able to mimic the fitness loss associated with resistance mutations by directly reducing the level of virion-associated protease activity. Virions containing 50% of a D25A mutant protease were 3- to 5-fold more sensitive to protease inhibitors. This level of reduction in protease activity also resulted in a 2-fold increase in sensitivity to nonnucleoside inhibitors of reverse transcriptase and a similar increase in sensitivity to zidovudine (AZT), indicating a pleiotropic effect associated with reduced protease activity. These results highlight the interplay between enzyme activity, viral fitness, and inhibitor mechanism and sensitivity in the closed system of the viral replication complex.
Figures




Similar articles
-
Cleavage of the murine leukemia virus transmembrane env protein by human immunodeficiency virus type 1 protease: transdominant inhibition by matrix mutations.J Virol. 1998 Dec;72(12):9621-7. doi: 10.1128/JVI.72.12.9621-9627.1998. J Virol. 1998. PMID: 9811695 Free PMC article.
-
Effect of mutations in the human immunodeficiency virus type 1 protease on cleavage of the gp41 cytoplasmic tail.J Virol. 2010 Mar;84(6):3121-6. doi: 10.1128/JVI.02002-09. Epub 2009 Dec 30. J Virol. 2010. PMID: 20042499 Free PMC article.
-
Nelfinavir-resistant, amprenavir-hypersusceptible strains of human immunodeficiency virus type 1 carrying an N88S mutation in protease have reduced infectivity, reduced replication capacity, and reduced fitness and process the Gag polyprotein precursor aberrantly.J Virol. 2002 Sep;76(17):8659-66. doi: 10.1128/jvi.76.17.8659-8666.2002. J Virol. 2002. PMID: 12163585 Free PMC article.
-
Structural and thermodynamic basis of resistance to HIV-1 protease inhibition: implications for inhibitor design.Curr Drug Targets Infect Disord. 2003 Dec;3(4):311-28. doi: 10.2174/1568005033481051. Curr Drug Targets Infect Disord. 2003. PMID: 14754432 Review.
-
Resistance to protease inhibitors.J HIV Ther. 2002 Nov;7(4):87-91. J HIV Ther. 2002. PMID: 12733606 Review.
Cited by
-
Direct-acting antiviral resistance of Hepatitis C virus is promoted by epistasis.Nat Commun. 2023 Nov 17;14(1):7457. doi: 10.1038/s41467-023-42550-6. Nat Commun. 2023. PMID: 37978179 Free PMC article.
-
Loss of the protease dimerization inhibition activity of tipranavir (TPV) and its association with the acquisition of resistance to TPV by HIV-1.J Virol. 2012 Dec;86(24):13384-96. doi: 10.1128/JVI.07234-11. Epub 2012 Sep 26. J Virol. 2012. PMID: 23015723 Free PMC article.
-
Mi3-GPU: MCMC-based Inverse Ising Inference on GPUs for protein covariation analysis.Comput Phys Commun. 2021 Mar;260:107312. doi: 10.1016/j.cpc.2020.107312. Epub 2020 Apr 17. Comput Phys Commun. 2021. PMID: 33716309 Free PMC article.
-
Novel P2 tris-tetrahydrofuran group in antiviral compound 1 (GRL-0519) fills the S2 binding pocket of selected mutants of HIV-1 protease.J Med Chem. 2013 Feb 14;56(3):1074-83. doi: 10.1021/jm301519z. Epub 2013 Jan 23. J Med Chem. 2013. PMID: 23298236 Free PMC article.
-
Treatment-associated polymorphisms in protease are significantly associated with higher viral load and lower CD4 count in newly diagnosed drug-naive HIV-1 infected patients.Retrovirology. 2012 Oct 3;9:81. doi: 10.1186/1742-4690-9-81. Retrovirology. 2012. PMID: 23031662 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

