Is the genetic landscape of the deep subsurface biosphere affected by viruses?
- PMID: 22084639
- PMCID: PMC3211056
- DOI: 10.3389/fmicb.2011.00219
Is the genetic landscape of the deep subsurface biosphere affected by viruses?
Abstract
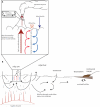
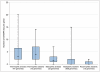
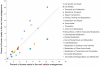
Viruses are powerful manipulators of microbial diversity, biogeochemistry, and evolution in the marine environment. Viruses can directly influence the genetic capabilities and the fitness of their hosts through the use of fitness factors and through horizontal gene transfer. However, the impact of viruses on microbial ecology and evolution is often overlooked in studies of the deep subsurface biosphere. Subsurface habitats connected to hydrothermal vent systems are characterized by constant fluid flux, dynamic environmental variability, and high microbial diversity. In such conditions, high adaptability would be an evolutionary asset, and the potential for frequent host-virus interactions would be high, increasing the likelihood that cellular hosts could acquire novel functions. Here, we review evidence supporting this hypothesis, including data indicating that microbial communities in subsurface hydrothermal fluids are exposed to a high rate of viral infection, as well as viral metagenomic data suggesting that the vent viral assemblage is particularly enriched in genes that facilitate horizontal gene transfer and host adaptability. Therefore, viruses are likely to play a crucial role in facilitating adaptability to the extreme conditions of these regions of the deep subsurface biosphere. We also discuss how these results might apply to other regions of the deep subsurface, where the nature of virus-host interactions would be altered, but possibly no less important, compared to more energetic hydrothermal systems.
Keywords: deep subsurface biosphere; hydrothermal vents; microbial evolution; viral ecology.
Figures


References
-
- Angly F. E., Felts B., Breitbart M., Salamon P., Edwards R. A., Carlson C., Chan A. M., Haynes M., Kelley S., Liu H., Mahaffy J. M., Mueller J. E., Nulton J., Olson R., Parsons R., Rayhawk S., Suttle C. A., Rohwer F. (2006). The marine viromes of four oceanic regions. PLoS Biol. 4, e368.10.1371/journal.pbio.0040368 - DOI - PMC - PubMed
-
- Baross J. A., Hoffman S. E. (1985). Submarine hydrothermal vents and associated gradient environments as sites for the origin and evolution of life. Orig. Life Evol. Biosph. 15, 327–34510.1007/BF01808177 - DOI
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

