Chloroviruses: not your everyday plant virus
- PMID: 22100667
- PMCID: PMC3259250
- DOI: 10.1016/j.tplants.2011.10.005
Chloroviruses: not your everyday plant virus
Abstract
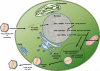
Viruses infecting higher plants are among the smallest viruses known and typically have four to ten protein-encoding genes. By contrast, many viruses that infect algae (classified in the virus family Phycodnaviridae) are among the largest viruses found to date and have up to 600 protein-encoding genes. This brief review focuses on one group of plaque-forming phycodnaviruses that infect unicellular chlorella-like green algae. The prototype chlorovirus PBCV-1 has more than 400 protein-encoding genes and 11 tRNA genes. About 40% of the PBCV-1 encoded proteins resemble proteins of known function including many that are completely unexpected for a virus. In many respects, chlorovirus infection resembles bacterial infection by tailed bacteriophages.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures


References
-
- Hull R. Comparative Plant Virology. Elsevier/Academic Press; Amsterdam: 2009.
-
- Nagasaki K. Dinoflagellates, diatoms, and their viruses. J. Microbiol. 2008;46:235–243. - PubMed
-
- Fuhrman JA. Marine viruses and their biogeochemical and ecological effects. Nature. 1999;399:541–548. - PubMed
-
- Suttle CA. Marine viruses - major players in the global ecosystem. Nat. Rev. Microbiol. 2005;5:801–812. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

