Substrate binding of a GH5 endoglucanase from the ruminal fungus Piromyces rhizinflata
- PMID: 22102024
- PMCID: PMC3212359
- DOI: 10.1107/S1744309111032428
Substrate binding of a GH5 endoglucanase from the ruminal fungus Piromyces rhizinflata
Abstract
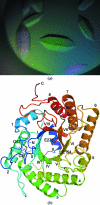
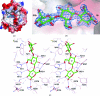
The endoglucanase EglA from Piromyces rhizinflata found in cattle stomach belongs to the GH5 family of glycoside hydrolases. The crystal structure of the catalytic domain of EglA shows the (β/α)(8)-barrel fold typical of GH5 enzymes. Adjacent to the active site of EglA, a loop containing a disulfide bond not found in other similar structures may participate in substrate binding. Because the active site was blocked by the N-terminal His tag of a neighbouring protein molecule in the crystal, enzyme-substrate complexes could not be obtained by soaking but were prepared by cocrystallization. The E154A mutant structure with a cellotriose bound to the -3, -2 and -1 subsites shows an extensive hydrogen-bonding network between the enzyme and the substrate, along with a stacking interaction between Trp44 and the -3 sugar. A possible dimer was observed in the crystal structure, but retention of activity in the E242A mutant suggested that the enzyme probably does not function as a dimer in solution. On the other hand, the first 100 amino acids encoded by the original cDNA fragment are very similar to those in the last third of the (β/α)(8)-barrel fold, indicating that EglA comprises at least two catalytic domains acting in tandem.
© 2011 International Union of Crystallography. All rights reserved.
Figures

Similar articles
-
Structure and enzymatic characterization of CelD endoglucanase from the anaerobic fungus Piromyces finnis.Appl Microbiol Biotechnol. 2023 Oct;107(19):5999-6011. doi: 10.1007/s00253-023-12684-0. Epub 2023 Aug 7. Appl Microbiol Biotechnol. 2023. PMID: 37548665 Free PMC article.
-
Immobilization of Piromyces rhizinflata β-glucanase on poly(dimethylsiloxane) and Si wafer and prediction of optimum reaction for enzyme activity.Prep Biochem Biotechnol. 2015;45(1):42-55. doi: 10.1080/10826068.2014.887579. Prep Biochem Biotechnol. 2015. PMID: 24670042
-
Primary sequence and enzymic properties of two modular endoglucanases, Cel5A and Cel45A, from the anaerobic fungus Piromyces equi.Microbiology (Reading). 2000 Aug;146 ( Pt 8):1999-2008. doi: 10.1099/00221287-146-8-1999. Microbiology (Reading). 2000. PMID: 10931904
-
The crystal structure of a family 5 endoglucanase mutant in complexed and uncomplexed forms reveals an induced fit activation mechanism.J Mol Biol. 1996 Apr 19;257(5):1042-51. doi: 10.1006/jmbi.1996.0222. J Mol Biol. 1996. PMID: 8632467
-
An intron-containing glycoside hydrolase family 9 cellulase gene encodes the dominant 90 kDa component of the cellulosome of the anaerobic fungus Piromyces sp. strain E2.Biochem J. 2002 Jul 1;365(Pt 1):193-204. doi: 10.1042/BJ20011866. Biochem J. 2002. PMID: 12071852 Free PMC article.
Cited by
-
A structural and kinetic survey of GH5_4 endoglucanases reveals determinants of broad substrate specificity and opportunities for biomass hydrolysis.J Biol Chem. 2020 Dec 18;295(51):17752-17769. doi: 10.1074/jbc.RA120.015328. J Biol Chem. 2020. PMID: 33454012 Free PMC article.
-
Characterization of two GH5 endoglucanases from termite microbiome using synthetic metagenomics.Appl Microbiol Biotechnol. 2020 Oct;104(19):8351-8366. doi: 10.1007/s00253-020-10831-5. Epub 2020 Aug 20. Appl Microbiol Biotechnol. 2020. PMID: 32816085
-
Structural Features of a Bacteroidetes-Affiliated Cellulase Linked with a Polysaccharide Utilization Locus.Sci Rep. 2015 Jul 2;5:11666. doi: 10.1038/srep11666. Sci Rep. 2015. PMID: 26133573 Free PMC article.
-
Carbohydrate Binding Modules: Diversity of Domain Architecture in Amylases and Cellulases From Filamentous Microorganisms.Front Bioeng Biotechnol. 2020 Jul 31;8:871. doi: 10.3389/fbioe.2020.00871. eCollection 2020. Front Bioeng Biotechnol. 2020. PMID: 32850729 Free PMC article. Review.
-
Activity and Thermostability of GH5 Endoglucanase Chimeras from Mesophilic and Thermophilic Parents.Appl Environ Microbiol. 2019 Feb 20;85(5):e02079-18. doi: 10.1128/AEM.02079-18. Print 2019 Mar 1. Appl Environ Microbiol. 2019. PMID: 30552196 Free PMC article.
References
-
- Bayer, E. A., Shimon, L. J., Shoham, Y. & Lamed, R. (1998). J. Struct. Biol. 124, 221–234. - PubMed
-
- Bhat, M. K. & Bhat, S. (1997). Biotechnol. Adv. 15, 583–620. - PubMed
-
- Bortoli-German, I., Haiech, J., Chippaux, M. & Barras, F. (1995). J. Mol. Biol. 246, 82–94. - PubMed
-
- Brünger, A. T., Adams, P. D., Clore, G. M., DeLano, W. L., Gros, P., Grosse-Kunstleve, R. W., Jiang, J.-S., Kuszewski, J., Nilges, M., Pannu, N. S., Read, R. J., Rice, L. M., Simonson, T. & Warren, G. L. (1998). Acta Cryst. D54, 905–921. - PubMed
-
- Collaborative Computational Project, Number 4 (1994). Acta Cryst. D50, 760–763. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources

