Sensitive RNA detection by combining three-way junction formation and primer generation-rolling circle amplification
- PMID: 22127872
- PMCID: PMC3273829
- DOI: 10.1093/nar/gkr909
Sensitive RNA detection by combining three-way junction formation and primer generation-rolling circle amplification
Abstract
Recently, we developed a simple isothermal nucleic acid amplification reaction, primer generation-rolling circle amplification (PG-RCA), to detect specific DNA sequences with great sensitivity and large dynamic range. In this paper, we combined PG-RCA with a three-way junction (3WJ) formation, and detected specific RNA molecules with high sensitivity and specificity in a one-step and isothermal reaction format. In the presence of target RNA, 3WJ probes (primer and template) are designed to form a 3WJ structure, from which multiple signal primers for the following PG-RCA can be generated by repeating primer extension, nicking and signal primer dissociation. Although this signal primer generation is a linear amplification process, the PG-RCA exponentially can amplify these signal primers and thus even a very small amount of RNA specimen can be detected. After optimizing the structures of 3WJ probes, the detection limit of this assay was 15.9 zmol (9.55 × 10(3) molecules) of synthetic RNA or 143 zmol (8.6 × 10(4) molecules) of in vitro transcribed human CD4 mRNA. Further, the applicability of this assay to detect CD4 mRNA in a human mRNA sample was demonstrated.
Figures



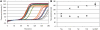



Similar articles
-
Sensitive isothermal detection of nucleic-acid sequence by primer generation-rolling circle amplification.Nucleic Acids Res. 2009 Feb;37(3):e19. doi: 10.1093/nar/gkn1014. Epub 2008 Dec 23. Nucleic Acids Res. 2009. PMID: 19106144 Free PMC article.
-
Urinary exosomal mRNA detection using novel isothermal gene amplification method based on three-way junction.Biosens Bioelectron. 2020 Nov 1;167:112474. doi: 10.1016/j.bios.2020.112474. Epub 2020 Aug 5. Biosens Bioelectron. 2020. PMID: 32798804
-
Sensitive fluorescent detection of DNA methyltransferase using nicking endonuclease-mediated multiple primers-like rolling circle amplification.Biosens Bioelectron. 2017 May 15;91:417-423. doi: 10.1016/j.bios.2016.12.061. Epub 2016 Dec 30. Biosens Bioelectron. 2017. PMID: 28063390
-
Recent advances in rolling circle amplification-based biosensing strategies-A review.Anal Chim Acta. 2021 Mar 1;1148:238187. doi: 10.1016/j.aca.2020.12.062. Epub 2020 Dec 31. Anal Chim Acta. 2021. PMID: 33516384 Review.
-
Rolling circle amplification: applications in nanotechnology and biodetection with functional nucleic acids.Angew Chem Int Ed Engl. 2008;47(34):6330-7. doi: 10.1002/anie.200705982. Angew Chem Int Ed Engl. 2008. PMID: 18680110 Review.
Cited by
-
Paper-based netlike rolling circle amplification (NRCA) for ultrasensitive and visual detection of SARS-CoV-2.Sens Actuators B Chem. 2022 May 1;358:131460. doi: 10.1016/j.snb.2022.131460. Epub 2022 Jan 22. Sens Actuators B Chem. 2022. PMID: 35095201 Free PMC article.
-
Production of dumbbell probe through hairpin cleavage-ligation and increasing RCA sensitivity and specificity by circle to circle amplification.Sci Rep. 2016 Jul 7;6:29229. doi: 10.1038/srep29229. Sci Rep. 2016. PMID: 27385060 Free PMC article.
-
Programmable High-Speed and Hyper-Efficiency DNA Signal Magnifier.Adv Sci (Weinh). 2022 Feb;9(4):e2104084. doi: 10.1002/advs.202104084. Epub 2021 Dec 16. Adv Sci (Weinh). 2022. PMID: 34913619 Free PMC article.
-
A short antisense oligonucleotide ameliorates symptoms of severe mouse models of spinal muscular atrophy.Mol Ther Nucleic Acids. 2014 Jul 8;3(7):e174. doi: 10.1038/mtna.2014.23. Mol Ther Nucleic Acids. 2014. PMID: 25004100 Free PMC article.
-
Detection of SARS-CoV-2 RNA through tandem isothermal gene amplification without reverse transcription.Anal Chim Acta. 2022 Jun 15;1212:339909. doi: 10.1016/j.aca.2022.339909. Epub 2022 May 6. Anal Chim Acta. 2022. PMID: 35623783 Free PMC article.
References
-
- Sambrook J, Russell DW. Molecular Cloning: A Laboratory Manual. 3rd edn. Harbor, NY: Cold Spring Harbor Laboratory Press, Cold Spring; 2001.
-
- Hod Y. A simplified ribonuclease protection assay. BioTechniques. 1992;13:852–854. - PubMed
-
- Saccomanno CF, Bordonaro M, Chen JS, Nordstrom JL. A faster ribonuclease protection assay. BioTechniques. 1992;13:846–850. - PubMed
-
- Bustin SA. Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. J. Mol. Endocrinol. 2000;25:169–193. - PubMed
-
- Schena M, Shalon D, Davis RW, Brown PO. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science. 1995;270:467–470. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

