Transposase mediated construction of RNA-seq libraries
- PMID: 22128135
- PMCID: PMC3246200
- DOI: 10.1101/gr.127373.111
Transposase mediated construction of RNA-seq libraries
Erratum in
- Genome Res. 2012 Mar;22(3):592
Abstract
RNA-seq has been widely adopted as a gene-expression measurement tool due to the detail, resolution, and sensitivity of transcript characterization that the technique provides. Here we present two transposon-based methods that efficiently construct high-quality RNA-seq libraries. We first describe a method that creates RNA-seq libraries for Illumina sequencing from double-stranded cDNA with only two enzymatic reactions. We generated high-quality RNA-seq libraries from as little as 10 pg of mRNA (∼1 ng of total RNA) with this approach. We also present a strand-specific RNA-seq library construction protocol that combines transposon-based library construction with uracil DNA glycosylase and endonuclease VIII to specifically degrade the second strand constructed during cDNA synthesis. The directional RNA-seq libraries maintain the same quality as the nondirectional libraries, while showing a high degree of strand specificity, such that 99.5% of reads map to the expected genomic strand. Each transposon-based library construction method performed well when compared with standard RNA-seq library construction methods with regard to complexity of the libraries, correlation between biological replicates, and the percentage of reads that align to the genome as well as exons. Our results show that high-quality RNA-seq libraries can be constructed efficiently and in an automatable fashion using transposition technology.
Figures

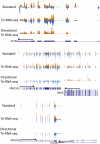



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
