Stem cell gene SALL4 suppresses transcription through recruitment of DNA methyltransferases
- PMID: 22128185
- PMCID: PMC3265879
- DOI: 10.1074/jbc.M111.308734
Stem cell gene SALL4 suppresses transcription through recruitment of DNA methyltransferases
Abstract
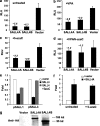
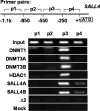
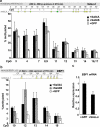
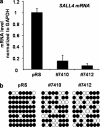
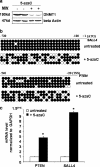
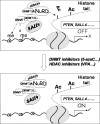
The stem cell protein SALL4 plays a vital role in maintaining stem cell identity and governing stem cell self-renewal through transcriptional repression. To explore SALL4-mediated mechanisms involved in transcriptional repression, we investigated DNA modifications underlying its regulatory activities. By a luciferase activity assay, we found that both histone deacetylase inhibitor valproic acid (VPA) and DNA methylation inhibitor 5-azacytidine (5-azaC) specifically reversed the repression effect of SALL4 on its own as well as other Sal gene promoter activities. Cotreatment of VPA with 5-azaC in cells almost completely blocked this repression effect. Further co-immunoprecipitation assay and enzyme activity analysis demonstrated that SALL4 protein directly interacted with different DNA methyltransferases (DNMTs) and purified DNMT enzymatic activities from nuclear extracts. In addition, SALL4 isoforms co-occupied the same regions of its own promoter as DNMT corepressors, and ectopic overexpression of SALL4 led to increased CpG island promoter methylation of silenced genes in various cell types. These included primary hematopoietic stem/progenitor cells, fibroblasts, and NB4 leukemic cells. In NB4 cells, treatment of cells with 5-azaC also caused decreased amounts of methylated alleles of SALL4 and PTEN and dramatically increased their mRNA expression. Our studies identify a new mechanism by which SALL4 represses gene expression through interaction with DNMTs. Furthermore, DNMTs and histone deacetylase repressors synergistically contribute to the regulatory effects of SALL4. These findings provide new insights into stem cell self-renewal mediated by SALL4 via epigenetic machinery.
Figures


Similar articles
-
Histone lysine-specific demethylase 1 (LSD1) protein is involved in Sal-like protein 4 (SALL4)-mediated transcriptional repression in hematopoietic stem cells.J Biol Chem. 2013 Nov 29;288(48):34719-28. doi: 10.1074/jbc.M113.506568. Epub 2013 Oct 25. J Biol Chem. 2013. PMID: 24163373 Free PMC article.
-
Precipitous release of methyl-CpG binding protein 2 and histone deacetylase 1 from the methylated human multidrug resistance gene (MDR1) on activation.Mol Cell Biol. 2002 Mar;22(6):1844-57. doi: 10.1128/MCB.22.6.1844-1857.2002. Mol Cell Biol. 2002. PMID: 11865062 Free PMC article.
-
Stem cell factor SALL4 represses the transcriptions of PTEN and SALL1 through an epigenetic repressor complex.PLoS One. 2009;4(5):e5577. doi: 10.1371/journal.pone.0005577. Epub 2009 May 18. PLoS One. 2009. PMID: 19440552 Free PMC article.
-
Reversal of gene silencing as a therapeutic target for cancer--roles for DNA methylation and its interdigitation with chromatin.Novartis Found Symp. 2004;259:226-33; discussion 234-7, 285-8. Novartis Found Symp. 2004. PMID: 15171257 Review.
-
Reactivation of silenced genes and transcriptional therapy.Cytogenet Genome Res. 2003;100(1-4):56-64. doi: 10.1159/000072838. Cytogenet Genome Res. 2003. PMID: 14526164 Review.
Cited by
-
Epigenetic Regulation of Hematopoietic Stem Cells.Int J Stem Cells. 2016 May 30;9(1):36-43. doi: 10.15283/ijsc.2016.9.1.36. Int J Stem Cells. 2016. PMID: 27426084 Free PMC article. Review.
-
Coordinating Regulation of Gene Expression in Cardiovascular Disease: Interactions between Chromatin Modifiers and Transcription Factors.Front Cardiovasc Med. 2017 Apr 6;4:19. doi: 10.3389/fcvm.2017.00019. eCollection 2017. Front Cardiovasc Med. 2017. PMID: 28428957 Free PMC article. Review.
-
Epigenetic silencing of SALL3 is an independent predictor of poor survival in head and neck cancer.Clin Epigenetics. 2017 Jun 12;9:64. doi: 10.1186/s13148-017-0363-1. eCollection 2017. Clin Epigenetics. 2017. PMID: 28616099 Free PMC article.
-
Upregulation of the Oct3/4 Network in Basal Breast Cancer Is Associated with Its Metastatic Potential and Shows Tissue Dependent Variability.Int J Mol Sci. 2023 May 23;24(11):9142. doi: 10.3390/ijms24119142. Int J Mol Sci. 2023. PMID: 37298091 Free PMC article.
-
Knockdown of SALL4 Protein Enhances All-trans Retinoic Acid-induced Cellular Differentiation in Acute Myeloid Leukemia Cells.J Biol Chem. 2015 Apr 24;290(17):10599-609. doi: 10.1074/jbc.M114.634790. Epub 2015 Mar 3. J Biol Chem. 2015. PMID: 25737450 Free PMC article.
References
-
- Kohlhase J., Schuh R., Dowe G., Kühnlein R. P., Jäckle H., Schroeder B., Schulz-Schaeffer W., Kretzschmar H. A., Köhler A., Müller U., Raab-Vetter M., Burkhardt E., Engel W., Stick R. (1996) Isolation, characterization, and organ-specific expression of two novel human zinc finger genes related to the Drosophila gene spalt. Genomics 38, 291–298 - PubMed
-
- Kühnlein R. P., Schuh R. (1996) Dual function of the region-specific homeotic gene spalt during Drosophila tracheal system development. Development 122, 2215–2223 - PubMed
-
- Al-Baradie R., Yamada K., St Hilaire C., Chan W. M., Andrews C., McIntosh N., Nakano M., Martonyi E. J., Raymond W. R., Okumura S., Okihiro M. M., Engle E. C. (2002) Duane radial ray syndrome (Okihiro syndrome) maps to 20q13 and results from mutations in SALL4, a new member of the SAL family. Am. J. Hum. Genet. 71, 1195–1199 - PMC - PubMed
-
- Kohlhase J., Heinrich M., Schubert L., Liebers M., Kispert A., Laccone F., Turnpenny P., Winter R. M., Reardon W. (2002) Okihiro syndrome is caused by SALL4 mutations. Hum. Mol. Genet. 11, 2979–2987 - PubMed
-
- Borozdin W., Wright M. J., Hennekam R. C., Hannibal M. C., Crow Y. J., Neumann T. E., Kohlhase J. (2004) Novel mutations in the gene SALL4 provide further evidence for acro-renal-ocular and Okihiro syndromes being allelic entities and extend the phenotypic spectrum. J. Med. Genet. 41, e102. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

