Expression of caveolin in trabecular meshwork cells and its possible implication in pathogenesis of primary open angle glaucoma
- PMID: 22128235
- PMCID: PMC3225292
Expression of caveolin in trabecular meshwork cells and its possible implication in pathogenesis of primary open angle glaucoma
Abstract
Purpose: Primary open-angle glaucoma (POAG), which is the most common form of glaucoma, has been associated with a heterogeneous genetic component. A genome-wide association study has identified a common sequence variant at 7q31 (rs4236601 [A]) near the caveolin genes in patients with POAG. Caveolins are a family of integral membrane proteins which participate in many cellular processes, including vesicular transport, cholesterol homeostasis, signal transduction, cell adhesion and migration. The goal of this study was to investigate the expression and regulation of caveolin 1 (CAV-1) and caveolin 2 (CAV-2) in normal and glaucoma trabecular meshwork (TM) cells.
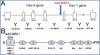
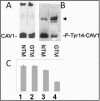
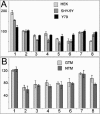
Methods: CAV-1 and CAV-2 protein expression was quantified by immunoblot analysis using lysates isolated from primary and immortalized TM cells or TM tissue dissected from normal and POAG eyes. The localization of caveolins in TM cells was assessed by immunofluorescent microscopy. CAV-1 and CAV-2 protein expression was also investigated in TM cells at various time points after subjecting the cells to known glaucomatous insults like dexamethasone (DEX) and tumor growth factor beta2 (TGF-β2) treatment. Phosphorylation of CAV-1 at tyrosine 14 in normal and glaucoma TM cell lines was evaluated using a specific monoclonal antibody (Ab). The 5' upstream region of the CAV-1 gene was amplified and the sequence variant rs4236601 (A/G polymorphic site) and several putative transcription factor-binding sites were modified by in vitro mutagenesis. The effect of nucleotide sequence modifications in the CAV-1 upstream region on gene expression was assayed in a luciferase-based system in TM and non-TM cells.
Results: CAV-1 and CAV-2 are expressed in TM cells, with localization to the cytoplasm and perinuclear region. DEX increased CAV-1 expression in immortalized glaucoma TM cells by 2.8±0.1 (n=3) fold at 24 h and 2.5±0.1 (n=3) fold at 48 h, compared to 1.3±0.06 (n=3) fold at 24 and 48 h in immortalized normal TM cells. Phosphorylation of CAV-1 at Tyr14 was reduced by 3.2±0.15 (n=3) fold in glaucomatous TM cells when compared to normal TM cells. In POAG and normal TM tissue, CAV-1 expression was found to be uniform. CAV-2, on the other hand, was variable in independent normal and glaucoma TM tissue. Substitution of a G for an A at base pair -2,388 upstream of the start codon of CAV-1, corresponding to the minor allele rs4236601 [A], increased transcriptional activity in TM and non-TM cells when compared to the native sequence. Deletion analysis of putative transcription factor binding sites in the CAV-1 promoter region caused cell-specific effects on gene expression.
Conclusions: CAV-1 and CAV-2 are expressed in normal and glaucoma tissue and TM cell lines. Phosphorylation of Tyr14 in CAV-1 and transcriptional regulation of CAV-1 expression may have a role in glaucomatous alterations in TM cells.
Figures



Similar articles
-
Regulation of glucocorticoid responsiveness in glaucomatous trabecular meshwork cells by glucocorticoid receptor-beta.Invest Ophthalmol Vis Sci. 2005 Dec;46(12):4607-16. doi: 10.1167/iovs.05-0571. Invest Ophthalmol Vis Sci. 2005. PMID: 16303956
-
Glaucomatous MYOC mutations activate the IL-1/NF-κB inflammatory stress response and the glaucoma marker SELE in trabecular meshwork cells.Mol Vis. 2015 Sep 17;21:1071-84. eCollection 2015. Mol Vis. 2015. PMID: 26396484 Free PMC article.
-
Effects of TGF-beta2, BMP-4, and gremlin in the trabecular meshwork: implications for glaucoma.Invest Ophthalmol Vis Sci. 2007 Mar;48(3):1191-200. doi: 10.1167/iovs.06-0296. Invest Ophthalmol Vis Sci. 2007. PMID: 17325163
-
The role of TGF-β2 and bone morphogenetic proteins in the trabecular meshwork and glaucoma.J Ocul Pharmacol Ther. 2014 Mar-Apr;30(2-3):154-62. doi: 10.1089/jop.2013.0220. Epub 2014 Feb 11. J Ocul Pharmacol Ther. 2014. PMID: 24517218 Free PMC article. Review.
-
Comprehensive bioinformatics analysis of trabecular meshwork gene expression data to unravel the molecular pathogenesis of primary open-angle glaucoma.Acta Ophthalmol. 2020 Feb;98(1):48-57. doi: 10.1111/aos.14154. Epub 2019 Jun 14. Acta Ophthalmol. 2020. PMID: 31197946 Free PMC article.
Cited by
-
Glaucoma Pathogenesis and Neurotrophins: Focus on the Molecular and Genetic Basis for Therapeutic Prospects.Curr Neuropharmacol. 2018;16(7):1018-1035. doi: 10.2174/1570159X16666180419121247. Curr Neuropharmacol. 2018. PMID: 29676228 Free PMC article. Review.
-
Activated αvβ3 integrin regulates αvβ5 integrin-mediated phagocytosis in trabecular meshwork cells.Invest Ophthalmol Vis Sci. 2013 Jul 24;54(7):5000-11. doi: 10.1167/iovs.13-12084. Invest Ophthalmol Vis Sci. 2013. PMID: 23821196 Free PMC article.
-
Differential effects of caveolin-1 and -2 knockdown on aqueous outflow and altered extracellular matrix turnover in caveolin-silenced trabecular meshwork cells.Invest Ophthalmol Vis Sci. 2014 Aug 7;55(9):5497-509. doi: 10.1167/iovs.14-14519. Invest Ophthalmol Vis Sci. 2014. PMID: 25103269 Free PMC article.
-
The landscape of GWAS validation; systematic review identifying 309 validated non-coding variants across 130 human diseases.BMC Med Genomics. 2022 Apr 1;15(1):74. doi: 10.1186/s12920-022-01216-w. BMC Med Genomics. 2022. PMID: 35365203 Free PMC article.
-
Intraocular pressure homeostasis: maintaining balance in a high-pressure environment.J Ocul Pharmacol Ther. 2014 Mar-Apr;30(2-3):94-101. doi: 10.1089/jop.2013.0185. Epub 2014 Jan 8. J Ocul Pharmacol Ther. 2014. PMID: 24401029 Free PMC article. Review.
References
-
- Quigley HA, Vitale S. Models of open-angle glaucoma prevalence and incidence in the United States. Invest Ophthalmol Vis Sci. 1997;38:83–91. - PubMed
-
- Thorleifsson G, Walters GB, Hewitt AW, Masson G, Helgason A, DeWan A, Sigurdsson A, Jonasdottir A, Gudjonsson SA, Magnusson KP, Stefansson H, Lam DS, Tam PO, Gudmundsdottir GJ, Southgate L, Burdon KP, Gottfredsdottir MS, Aldred MA, Mitchell P, St Clair D, Collier DA, Tang N, Sveinsson O, Macgregor S, Martin NG, Cree AJ, Gibson J, Macleod A, Jacob A, Ennis S, Young TL, Chan JC, Karwatowski WS, Hammond CJ, Thordarson K, Zhang M, Wadelius C, Lotery AJ, Trembath RC, Pang CP, Hoh J, Craig JE, Kong A, Mackey DA, Jonasson F, Thorsteinsdottir U, Stefansson K. Common variants near CAV-1 and CAV-2 are associated with primary open-angle glaucoma. Nat Genet. 2010;42:906–9. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
