Phylogenomics of Reichenowia parasitica, an alphaproteobacterial endosymbiont of the freshwater leech Placobdella parasitica
- PMID: 22132238
- PMCID: PMC3223239
- DOI: 10.1371/journal.pone.0028192
Phylogenomics of Reichenowia parasitica, an alphaproteobacterial endosymbiont of the freshwater leech Placobdella parasitica
Abstract
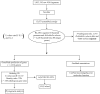
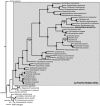
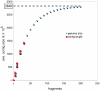
Although several commensal alphaproteobacteria form close relationships with plant hosts where they aid in (e.g.,) nitrogen fixation and nodulation, only a few inhabit animal hosts. Among these, Reichenowia picta, R. ornata and R. parasitica, are currently the only known mutualistic, alphaproteobacterial endosymbionts to inhabit leeches. These bacteria are harbored in the epithelial cells of the mycetomal structures of their freshwater leech hosts, Placobdella spp., and these structures have no other obvious function than housing bacterial symbionts. However, the function of the bacterial symbionts has remained unclear. Here, we focused both on exploring the genomic makeup of R. parasitica and on performing a robust phylogenetic analysis, based on more data than previous hypotheses, to test its position among related bacteria. We sequenced a combined pool of host and symbiont DNA from 36 pairs of mycetomes and performed an in silico separation of the different DNA pools through subtractive scaffolding. The bacterial contigs were compared to 50 annotated bacterial genomes and the genome of the freshwater leech Helobdella robusta using a BLASTn protocol. Further, amino acid sequences inferred from the contigs were used as queries against the 50 bacterial genomes to establish orthology. A total of 358 orthologous genes were used for the phylogenetic analyses. In part, results suggest that R. parasitica possesses genes coding for proteins related to nitrogen fixation, iron/vitamin B translocation and plasmid survival. Our results also indicate that R. parasitica interacts with its host in part by transmembrane signaling and that several of its genes show orthology across Rhizobiaceae. The phylogenetic analyses support the nesting of R. parasitica within the Rhizobiaceae, as sister to a group containing Agrobacterium and Rhizobium species.
Conflict of interest statement
Figures


Similar articles
-
Leech mycetome endosymbionts are a new lineage of alphaproteobacteria related to the Rhizobiaceae.Mol Phylogenet Evol. 2004 Jan;30(1):178-86. doi: 10.1016/s1055-7903(03)00184-2. Mol Phylogenet Evol. 2004. PMID: 15022768
-
New gammaproteobacteria associated with blood-feeding leeches and a broad phylogenetic analysis of leech endosymbionts.Appl Environ Microbiol. 2005 Sep;71(9):5219-24. doi: 10.1128/AEM.71.9.5219-5224.2005. Appl Environ Microbiol. 2005. PMID: 16151107 Free PMC article.
-
Discovery of a novel symbiotic lineage associated with a hematophagous leech from the genus Haementeria.Microbiol Spectr. 2024 Jul 2;12(7):e0428623. doi: 10.1128/spectrum.04286-23. Epub 2024 Jun 6. Microbiol Spectr. 2024. PMID: 38842327 Free PMC article.
-
Symbiosis within Symbiosis: Evolving Nitrogen-Fixing Legume Symbionts.Trends Microbiol. 2016 Jan;24(1):63-75. doi: 10.1016/j.tim.2015.10.007. Epub 2015 Nov 21. Trends Microbiol. 2016. PMID: 26612499 Review.
-
Leeches and their microbiota: naturally simple symbiosis models.Trends Microbiol. 2006 Aug;14(8):365-71. doi: 10.1016/j.tim.2006.06.009. Epub 2006 Jul 14. Trends Microbiol. 2006. PMID: 16843660 Review.
Cited by
-
Solving a Bloody Mess: B-Vitamin Independent Metabolic Convergence among Gammaproteobacterial Obligate Endosymbionts from Blood-Feeding Arthropods and the Leech Haementeria officinalis.Genome Biol Evol. 2015 Oct 9;7(10):2871-84. doi: 10.1093/gbe/evv188. Genome Biol Evol. 2015. PMID: 26454017 Free PMC article.
-
Comparative Mitogenomics of Leeches (Annelida: Clitellata): Genome Conservation and Placobdella-Specific trnD Gene Duplication.PLoS One. 2016 May 13;11(5):e0155441. doi: 10.1371/journal.pone.0155441. eCollection 2016. PLoS One. 2016. PMID: 27176910 Free PMC article.
-
Ideating iDNA: Lessons and limitations from leeches in legacy collections.PLoS One. 2019 Feb 22;14(2):e0212226. doi: 10.1371/journal.pone.0212226. eCollection 2019. PLoS One. 2019. PMID: 30794582 Free PMC article.
References
-
- Graf J, Kikuchi Y, Rio RVM. Leeches and their microbiota: naturally simple symbiosis models. Trends Microbiol. 2006;14:365–371. - PubMed
-
- Reichenow E. Über intrazelluläre Symbionten bei Blutsaugern. Arch Schiffs-u Tropen-Hyg. 1921;25
-
- Reichenow E. Intrazelluläre Symbionten bei blutsaugenden Milben und Egeln Arch Protistenk. 1922;45
-
- Siddall ME, Perkins SL, Desser SS. Leech mycetome endosymbionts are a new lineage of alphaproteobacteria related to the Rhizobiaceae. Mol Phylogenet Evol. 2004;30:178–186. - PubMed
-
- Akman L, Yamashita A, Watanabe H, Oshima K, Shiba T, et al. Genome sequence of the endocellular obligate symbiont of tsetse flies, Wigglesworthia glossinida. Nat Genet. 2002;32:402–407. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

