Infrared study of the stability and folding kinetics of a series of β-hairpin peptides with a common NPDG turn
- PMID: 22136248
- PMCID: PMC3278157
- DOI: 10.1021/jp2046867
Infrared study of the stability and folding kinetics of a series of β-hairpin peptides with a common NPDG turn
Abstract
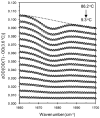
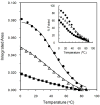
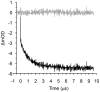
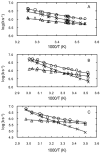
The thermal stability and folding kinetics of a series of 15-residue β-hairpins with a common Type I [3:5] NPDG turn were studied using Fourier transform infrared spectroscopy (FTIR) and laser-induced temperature jump (T-jump) with infrared detection, respectively. Mutations at positions 3, 5, or 13 in the peptide sequence SEXYXNPDGTWTXTE, where X represents the position of mutation, were performed to study the roles of hydrophobic interactions in determining the thermodynamic and kinetic properties of β-hairpin folding. The thermal stability studies show a broad thermal folding/unfolding transition for all the peptides. T-jump studies indicate that these β-hairpin peptides fold in less than 2 μs. In addition, both folding and unfolding rate constants decrease with increasing strength of hydrophobic interactions. Kinetically, the hydrophobic interactions have more significant influence on the unfolding rate than the folding rate. Φ-value analysis indicates that the hydrophobic interactions between the side chains are mainly formed at the latter part of the transition-state region during the folding process. In summary, the results suggest that the formation of the native structure of these β-hairpins depends on the correct topology of the hydrophobic cluster. Besides the formation of the turn region as a key process for folding as suggested by previous studies, a hydrophobic collapse process may also play a crucial role during β-hairpin folding.
© 2011 American Chemical Society
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

