A machine learning approach to identify hydrogenosomal proteins in Trichomonas vaginalis
- PMID: 22140228
- PMCID: PMC3272890
- DOI: 10.1128/EC.05225-11
A machine learning approach to identify hydrogenosomal proteins in Trichomonas vaginalis
Abstract
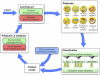
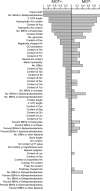
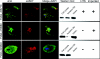
The protozoan parasite Trichomonas vaginalis is the causative agent of trichomoniasis, the most widespread nonviral sexually transmitted disease in humans. It possesses hydrogenosomes-anaerobic mitochondria that generate H(2), CO(2), and acetate from pyruvate while converting ADP to ATP via substrate-level phosphorylation. T. vaginalis hydrogenosomes lack a genome and translation machinery; hence, they import all their proteins from the cytosol. To date, however, only 30 imported proteins have been shown to localize to the organelle. A total of 226 nuclear-encoded proteins inferred from the genome sequence harbor a characteristic short N-terminal presequence, reminiscent of mitochondrial targeting peptides, which is thought to mediate hydrogenosomal targeting. Recent studies suggest, however, that the presequences might be less important than previously thought. We sought to identify new hydrogenosomal proteins within the 59,672 annotated open reading frames (ORFs) of T. vaginalis, independent of the N-terminal targeting signal, using a machine learning approach. Our training set included 57 gene and protein features determined for all 30 known hydrogenosomal proteins and 576 nonhydrogenosomal proteins. Several classifiers were trained on this set to yield an import score for all proteins encoded by T. vaginalis ORFs, predicting the likelihood of hydrogenosomal localization. The machine learning results were tested through immunofluorescence assay and immunodetection in isolated cell fractions of 14 protein predictions using hemagglutinin constructs expressed under the homologous SCSα promoter in transiently transformed T. vaginalis cells. Localization of 6 of the 10 top predicted hydrogenosome-localized proteins was confirmed, and two of these were found to lack an obvious N-terminal targeting signal.
Figures



Similar articles
-
Protein import into hydrogenosomes of Trichomonas vaginalis involves both N-terminal and internal targeting signals: a case study of thioredoxin reductases.Eukaryot Cell. 2008 Oct;7(10):1750-7. doi: 10.1128/EC.00206-08. Epub 2008 Aug 1. Eukaryot Cell. 2008. PMID: 18676956 Free PMC article.
-
The N-terminal sequences of four major hydrogenosomal proteins are not essential for import into hydrogenosomes of Trichomonas vaginalis.J Eukaryot Microbiol. 2013 Jan-Feb;60(1):89-97. doi: 10.1111/jeu.12012. Epub 2012 Dec 4. J Eukaryot Microbiol. 2013. PMID: 23210891
-
Competition and protease sensitivity assays provide evidence for the existence of a hydrogenosomal protein import machinery in Trichomonas vaginalis.Mol Biochem Parasitol. 2000 Feb 25;106(1):11-20. doi: 10.1016/s0166-6851(99)00196-6. Mol Biochem Parasitol. 2000. PMID: 10743607
-
Biogenesis of the hydrogenosome in the anaerobic protist Trichomonas vaginalis.J Parasitol. 1993 Oct;79(5):664-70. J Parasitol. 1993. PMID: 8410536 Review.
-
The hydrogenosome of Trichomonas vaginalis.J Eukaryot Microbiol. 2022 Nov;69(6):e12922. doi: 10.1111/jeu.12922. Epub 2022 May 30. J Eukaryot Microbiol. 2022. PMID: 35567536 Review.
Cited by
-
Evolution of myxozoan mitochondrial genomes: insights from myxobolids.BMC Genomics. 2024 Apr 22;25(1):388. doi: 10.1186/s12864-024-10254-w. BMC Genomics. 2024. PMID: 38649808 Free PMC article.
-
Reliability of plastid and mitochondrial localisation prediction declines rapidly with the evolutionary distance to the training set increasing.PLoS Comput Biol. 2024 Nov 11;20(11):e1012575. doi: 10.1371/journal.pcbi.1012575. eCollection 2024 Nov. PLoS Comput Biol. 2024. PMID: 39527633 Free PMC article.
-
Recent Advances in the Trichomonas vaginalis Field.F1000Res. 2016 Feb 11;5:F1000 Faculty Rev-162. doi: 10.12688/f1000research.7594.1. eCollection 2016. F1000Res. 2016. PMID: 26918168 Free PMC article. Review.
-
N-Terminal Presequence-Independent Import of Phosphofructokinase into Hydrogenosomes of Trichomonas vaginalis.Eukaryot Cell. 2015 Dec;14(12):1264-75. doi: 10.1128/EC.00104-15. Epub 2015 Oct 16. Eukaryot Cell. 2015. PMID: 26475173 Free PMC article.
-
A mitochondrion-free eukaryote contains proteins capable of import into an exogenous mitochondrion-related organelle.Open Biol. 2023 Jan;13(1):220238. doi: 10.1098/rsob.220238. Epub 2023 Jan 11. Open Biol. 2023. PMID: 36629021 Free PMC article.
References
-
- Akhmanova A, et al. 1998. A hydrogenosome with a genome. Nature 396:527–528 - PubMed
-
- Alsmark UC, Sicheritz-Ponten T, Foster PG, Hirt RP, Embley TM. 2009. Horizontal gene transfer in eukaryotic parasites: a case study of Entamoeba histolytica and Trichomonas vaginalis. Methods Mol. Biol. 532:489–500 - PubMed
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403–410 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

