On the evolutionary origin of eukaryotic DNA methyltransferases and Dnmt2
- PMID: 22140515
- PMCID: PMC3227630
- DOI: 10.1371/journal.pone.0028104
On the evolutionary origin of eukaryotic DNA methyltransferases and Dnmt2
Abstract
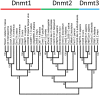
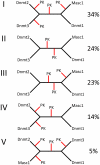
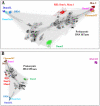
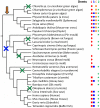
The Dnmt2 enzymes show strong amino acid sequence similarity with eukaryotic and prokaryotic DNA-(cytosine C5)-methyltransferases. Yet, Dnmt2 enzymes from several species were shown to methylate tRNA-Asp and had been proposed that eukaryotic DNA methyltransferases evolved from a Dnmt2-like tRNA methyltransferase ancestor [Goll et al., 2006, Science, 311, 395-8]. It was the aim of this study to investigate if this hypothesis could be supported by evidence from sequence alignments. We present phylogenetic analyses based on sequence alignments of the methyltransferase catalytic domains of more than 2300 eukaryotic and prokaryotic DNA-(cytosine C5)-methyltransferases and analyzed the distribution of DNA methyltransferases in eukaryotic species. The Dnmt2 homologues were reliably identified by an additional conserved CFT motif next to motif IX. All DNA methyltransferases and Dnmt2 enzymes were clearly separated from other RNA-(cytosine-C5)-methyltransferases. Our sequence alignments and phylogenetic analyses indicate that the last universal eukaryotic ancestor contained at least one member of the Dnmt1, Dnmt2 and Dnmt3 families of enzymes and additional RNA methyltransferases. The similarity of Dnmt2 enzymes with DNA methyltransferases and absence of similarity with RNA methyltransferases combined with their strong RNA methylation activity suggest that the ancestor of Dnmt2 was a DNA methyltransferase and an early Dnmt2 enzyme changed its substrate preference to tRNA. There is no phylogenetic evidence that Dnmt2 was the precursor of eukaryotic Dnmts. Most likely, the eukaryotic Dnmt1 and Dnmt3 families of DNA methyltransferases had an independent origin in the prokaryotic DNA methyltransferase sequence space.
Conflict of interest statement
Figures


Similar articles
-
Human DNMT2 methylates tRNA(Asp) molecules using a DNA methyltransferase-like catalytic mechanism.RNA. 2008 Aug;14(8):1663-70. doi: 10.1261/rna.970408. Epub 2008 Jun 20. RNA. 2008. PMID: 18567810 Free PMC article.
-
[Dnmt2 is the Most Evolutionary Conserved and Enigmatic Cytosine DNA Methyltransferase in Eukaryotes].Genetika. 2016 Mar;52(3):269-82. Genetika. 2016. PMID: 27281847 Review. Russian.
-
Solving the Dnmt2 enigma.Chromosoma. 2010 Feb;119(1):35-40. doi: 10.1007/s00412-009-0240-6. Chromosoma. 2010. PMID: 19730874 Review.
-
A candidate mammalian DNA methyltransferase related to pmt1p of fission yeast.Hum Mol Genet. 1998 Feb;7(2):279-84. doi: 10.1093/hmg/7.2.279. Hum Mol Genet. 1998. PMID: 9425235
-
Structure of human DNMT2, an enigmatic DNA methyltransferase homolog that displays denaturant-resistant binding to DNA.Nucleic Acids Res. 2001 Jan 15;29(2):439-48. doi: 10.1093/nar/29.2.439. Nucleic Acids Res. 2001. PMID: 11139614 Free PMC article.
Cited by
-
Epigenetic regulation of development and pathogenesis in fungal plant pathogens.Mol Plant Pathol. 2017 Aug;18(6):887-898. doi: 10.1111/mpp.12499. Epub 2016 Nov 28. Mol Plant Pathol. 2017. PMID: 27749982 Free PMC article. Review.
-
A map of 5-methylcytosine residues in Trypanosoma brucei tRNA revealed by sodium bisulfite sequencing.Mol Biochem Parasitol. 2014 Feb;193(2):122-6. doi: 10.1016/j.molbiopara.2013.12.003. Epub 2014 Jan 2. Mol Biochem Parasitol. 2014. PMID: 24389163 Free PMC article.
-
Network-based visualisation reveals new insights into transposable element diversity.Mol Syst Biol. 2021 Jun;17(6):e9600. doi: 10.15252/msb.20209600. Mol Syst Biol. 2021. PMID: 34169647 Free PMC article.
-
Reviving the RNA World: An Insight into the Appearance of RNA Methyltransferases.Front Genet. 2016 Jun 6;7:99. doi: 10.3389/fgene.2016.00099. eCollection 2016. Front Genet. 2016. PMID: 27375676 Free PMC article. Review.
-
DNA Methylation in Basal Metazoans: Insights from Ctenophores.Integr Comp Biol. 2015 Dec;55(6):1096-110. doi: 10.1093/icb/icv086. Epub 2015 Jul 14. Integr Comp Biol. 2015. PMID: 26173712 Free PMC article. Review.
References
-
- Klose RJ, Bird AP. Genomic DNA methylation: the mark and its mediators. Trends Biochem Sci. 2006;31:89–97. - PubMed
-
- Jurkowska RZ, Jurkowski TP, Jeltsch A. Structure and function of mammalian DNA methyltransferases. Chembiochem. 2011;12:206–222. - PubMed
-
- Goll MG, Bestor TH. Eukaryotic cytosine methyltransferases. Annu Rev Biochem. 2005;74:481–514. - PubMed
-
- Zemach A, McDaniel IE, Silva P, Zilberman D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science. 2010;328:916–919. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

