A systematic analysis on DNA methylation and the expression of both mRNA and microRNA in bladder cancer
- PMID: 22140553
- PMCID: PMC3227661
- DOI: 10.1371/journal.pone.0028223
A systematic analysis on DNA methylation and the expression of both mRNA and microRNA in bladder cancer
Abstract
Background: DNA methylation aberration and microRNA (miRNA) deregulation have been observed in many types of cancers. A systematic study of methylome and transcriptome in bladder urothelial carcinoma has never been reported.
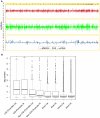
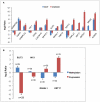
Methodology/principal findings: The DNA methylation was profiled by modified methylation-specific digital karyotyping (MMSDK) and the expression of mRNAs and miRNAs was analyzed by digital gene expression (DGE) sequencing in tumors and matched normal adjacent tissues obtained from 9 bladder urothelial carcinoma patients. We found that a set of significantly enriched pathways disrupted in bladder urothelial carcinoma primarily related to "neurogenesis" and "cell differentiation" by integrated analysis of -omics data. Furthermore, we identified an intriguing collection of cancer-related genes that were deregulated at the levels of DNA methylation and mRNA expression, and we validated several of these genes (HIC1, SLIT2, RASAL1, and KRT17) by Bisulfite Sequencing PCR and Reverse Transcription qPCR in a panel of 33 bladder cancer samples.
Conclusions/significance: We characterized the profiles between methylome and transcriptome in bladder urothelial carcinoma, identified a set of significantly enriched key pathways, and screened four aberrantly methylated and expressed genes. Conclusively, our findings shed light on a new avenue for basic bladder cancer research.
Conflict of interest statement
Figures
Similar articles
-
MicroRNA expression signatures of bladder cancer revealed by deep sequencing.PLoS One. 2011 Mar 28;6(3):e18286. doi: 10.1371/journal.pone.0018286. PLoS One. 2011. PMID: 21464941 Free PMC article.
-
Targeted DNA and RNA Sequencing of Paired Urothelial and Squamous Bladder Cancers Reveals Discordant Genomic and Transcriptomic Events and Unique Therapeutic Implications.Eur Urol. 2018 Dec;74(6):741-753. doi: 10.1016/j.eururo.2018.06.047. Epub 2018 Jul 20. Eur Urol. 2018. PMID: 30033047
-
A new algorithm for integrated analysis of miRNA-mRNA interactions based on individual classification reveals insights into bladder cancer.PLoS One. 2013 May 24;8(5):e64543. doi: 10.1371/journal.pone.0064543. Print 2013. PLoS One. 2013. PMID: 23717626 Free PMC article.
-
Predicting favourable prognosis of urothelial carcinoma: gene expression and genome profiling.Curr Opin Urol. 2009 Sep;19(5):516-21. doi: 10.1097/MOU.0b013e32832eb45f. Curr Opin Urol. 2009. PMID: 19553819 Review.
-
Alterations of DNA methylome in human bladder cancer.Epigenetics. 2013 Oct;8(10):1013-22. doi: 10.4161/epi.25927. Epub 2013 Aug 6. Epigenetics. 2013. PMID: 23975266 Free PMC article. Review.
Cited by
-
Cross-contamination of a UROtsa stock with T24 cells--molecular comparison of different cell lines and stocks.PLoS One. 2013 May 17;8(5):e64139. doi: 10.1371/journal.pone.0064139. Print 2013. PLoS One. 2013. PMID: 23691160 Free PMC article.
-
Differential mRNA expression profiling of oral squamous cell carcinoma by high-throughput RNA sequencing.J Biomed Res. 2015 Jan 30;29(5):397-404. doi: 10.7555/JBR.29.20140088. Online ahead of print. J Biomed Res. 2015. PMID: 26273018 Free PMC article.
-
Epigenetics of Bladder Cancer: Where Biomarkers and Therapeutic Targets Meet.Front Genet. 2019 Nov 18;10:1125. doi: 10.3389/fgene.2019.01125. eCollection 2019. Front Genet. 2019. PMID: 31850055 Free PMC article. Review.
-
Syndromic male subfertility: A network view of genome-phenome associations.Andrology. 2022 May;10(4):720-732. doi: 10.1111/andr.13167. Epub 2022 Mar 15. Andrology. 2022. PMID: 35218153 Free PMC article.
-
Identification of γ-synuclein as a stage-specific marker in bladder cancer by immunohistochemistry.Med Sci Monit. 2014 Dec 5;20:2550-5. doi: 10.12659/MSM.892927. Med Sci Monit. 2014. PMID: 25479371 Free PMC article.
References
-
- Sugimura T, Ushijima T. Genetic and epigenetic alterations in carcinogenesis. Mutat Res. 2000;462:235–246. - PubMed
-
- Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev Genet. 2002;3:415–428. - PubMed
-
- Eden A, Gaudet F, Waghmare A, Jaenisch R. Chromosomal instability and tumors promoted by DNA hypomethylation. Science. 2003;300:455. - PubMed
-
- Clark SJ, Melki J. DNA methylation and gene silencing in cancer: which is the guilty party? Oncogene. 2002;21:5380–5387. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

