Identification of a genomic reservoir for new TRIM genes in primate genomes
- PMID: 22144910
- PMCID: PMC3228819
- DOI: 10.1371/journal.pgen.1002388
Identification of a genomic reservoir for new TRIM genes in primate genomes
Abstract
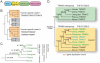
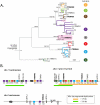
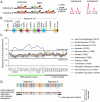
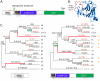
Tripartite Motif (TRIM) ubiquitin ligases act in the innate immune response against viruses. One of the best characterized members of this family, TRIM5α, serves as a potent retroviral restriction factor with activity against HIV. Here, we characterize what are likely to be the youngest TRIM genes in the human genome. For instance, we have identified 11 TRIM genes that are specific to humans and African apes (chimpanzees, bonobos, and gorillas) and another 7 that are human-specific. Many of these young genes have never been described, and their identification brings the total number of known human TRIM genes to approximately 100. These genes were acquired through segmental duplications, most of which originated from a single locus on chromosome 11. Another polymorphic duplication of this locus has resulted in these genes being copy number variable within the human population, with a Han Chinese woman identified as having 12 additional copies of these TRIM genes compared to other individuals screened in this study. Recently, this locus was annotated as one of 34 "hotspot" regions that are also copy number variable in the genomes of chimpanzees and rhesus macaques. Most of the young TRIM genes originating from this locus are expressed, spliced, and contain signatures of positive natural selection in regions known to determine virus recognition in TRIM5α. However, we find that they do not restrict the same retroviruses as TRIM5α, consistent with the high degree of divergence observed in the regions that control target specificity. We propose that this recombinationally volatile locus serves as a reservoir from which new TRIM genes arise through segmental duplication, allowing primates to continually acquire new antiviral genes that can be selected to target new and evolving pathogens.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

References
-
- McNab FW, Rajsbaum R, Stoye JP, O'Garra A. Tripartite-motif proteins and innate immune regulation. Curr Opin Immunol. 2011;23:46–56. - PubMed
-
- Nisole S, Stoye JP, Saïb A. TRIM family proteins: retroviral restriction and antiviral defence. Nat Rev Micro. 2005;3:799–808. - PubMed
-
- Stremlau M, Owens CM, Perron MJ, Kiessling M, Autissier P, et al. The cytoplasmic body component TRIM5alpha restricts HIV-1 infection in Old World monkeys. Nature. 2004;427:848–853. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

