The impact of phosphate scarcity on pharmaceutical protein production in S. cerevisiae: linking transcriptomic insights to phenotypic responses
- PMID: 22151908
- PMCID: PMC3265430
- DOI: 10.1186/1475-2859-10-104
The impact of phosphate scarcity on pharmaceutical protein production in S. cerevisiae: linking transcriptomic insights to phenotypic responses
Abstract
Background: The adaptation of unicellular organisms like Saccharomyces cerevisiae to alternating nutrient availability is of great fundamental and applied interest, as understanding how eukaryotic cells respond to variations in their nutrient supply has implications spanning from physiological insights to biotechnological applications.
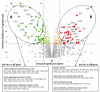
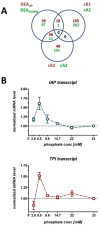
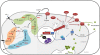
Results: The impact of a step-wise restricted supply of phosphate on the physiological state of S. cerevisiae cells producing human Insulin was studied. The focus was to determine the changes within the global gene expression of cells being cultured to an industrially relevant high cell density of 33 g/l cell dry weight and under six distinct phosphate concentrations, ranging from 33 mM (unlimited) to 2.6 mM (limited). An increased flux through the secretory pathway, being induced by the PHO circuit during low P(i) supplementation, proved to enhance the secretory production of the heterologous protein. The re-distribution of the carbon flux from biomass formation towards increased glycerol production under low phosphate led to increased transcript levels of the insulin gene, which was under the regulation of the TPI1 promoter.
Conclusions: Our study underlines the dynamic character of adaptive responses of cells towards a change in their nutrient access. The gradual decrease of the phosphate supply resulted in a step-wise modulated phenotypic response, thereby alternating the specific productivity and the secretory flux. Our work emphasizes the importance of reduced phosphate supply for improved secretory production of heterologous proteins.
Figures


Similar articles
-
Nutrient-regulated antisense and intragenic RNAs modulate a signal transduction pathway in yeast.PLoS Biol. 2008 Dec 23;6(12):2817-30. doi: 10.1371/journal.pbio.0060326. PLoS Biol. 2008. PMID: 19108609 Free PMC article.
-
New aspects on phosphate sensing and signalling in Saccharomyces cerevisiae.FEMS Yeast Res. 2006 Mar;6(2):171-6. doi: 10.1111/j.1567-1364.2006.00036.x. FEMS Yeast Res. 2006. PMID: 16487340 Review.
-
The competitive advantage of a dual-transporter system.Science. 2011 Dec 9;334(6061):1408-12. doi: 10.1126/science.1207154. Science. 2011. PMID: 22158820 Free PMC article.
-
Metabolic engineering for high glycerol production by the anaerobic cultures of Saccharomyces cerevisiae.Appl Microbiol Biotechnol. 2017 Jun;101(11):4403-4416. doi: 10.1007/s00253-017-8202-z. Epub 2017 Mar 9. Appl Microbiol Biotechnol. 2017. PMID: 28280870
-
Inorganic Phosphate and Sulfate Transport in S. cerevisiae.Adv Exp Med Biol. 2016;892:253-269. doi: 10.1007/978-3-319-25304-6_10. Adv Exp Med Biol. 2016. PMID: 26721277 Review.
Cited by
-
Large-scale production of tauroursodeoxycholic acid products through fermentation optimization of engineered Escherichia coli cell factory.Microb Cell Fact. 2019 Feb 8;18(1):34. doi: 10.1186/s12934-019-1076-2. Microb Cell Fact. 2019. PMID: 30736766 Free PMC article.
-
Physiological and proteomic analysis of Escherichia coli iron-limited chemostat growth.J Bacteriol. 2014 Aug;196(15):2748-61. doi: 10.1128/JB.01606-14. Epub 2014 May 16. J Bacteriol. 2014. PMID: 24837288 Free PMC article.
-
Cycles, sources, and sinks: Conceptualizing how phosphate balance modulates carbon flux using yeast metabolic networks.Elife. 2021 Feb 5;10:e63341. doi: 10.7554/eLife.63341. Elife. 2021. PMID: 33544078 Free PMC article. Review.
-
Design and Implementation of an Automated Illuminating, Culturing, and Sampling System for Microbial Optogenetic Applications.J Vis Exp. 2017 Feb 19;(120):54894. doi: 10.3791/54894. J Vis Exp. 2017. PMID: 28287505 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

