Structural organization of the kinetochore-microtubule interface
- PMID: 22154944
- PMCID: PMC3294040
- DOI: 10.1016/j.ceb.2011.11.003
Structural organization of the kinetochore-microtubule interface
Abstract
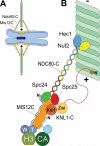
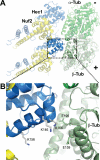
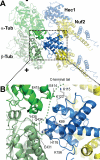
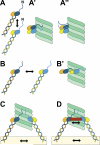
Successful mitosis depends on the stable, yet regulated attachment of chromosomes to spindle microtubules. The kinetochore, a large macromolecular structure assembled at sites of centromeric heterochromatin, is responsible for generating and regulating these essential attachments. Over the last several years, concerted experimental efforts have brought the structural view of the kinetochore-microtubule interface more clearly into focus. Here, we review important recent advancements and discuss several unresolved questions regarding how kinetochores dynamically bridge mitotic chromosomes to spindle microtubules.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures
Similar articles
-
CLIP-170 facilitates the formation of kinetochore-microtubule attachments.EMBO J. 2006 Jan 11;25(1):45-57. doi: 10.1038/sj.emboj.7600916. Epub 2005 Dec 15. EMBO J. 2006. PMID: 16362039 Free PMC article.
-
MAST/Orbit has a role in microtubule-kinetochore attachment and is essential for chromosome alignment and maintenance of spindle bipolarity.J Cell Biol. 2002 May 27;157(5):749-60. doi: 10.1083/jcb.200201101. Epub 2002 May 28. J Cell Biol. 2002. PMID: 12034769 Free PMC article.
-
Kinetochore flexibility: creating a dynamic chromosome-spindle interface.Curr Opin Cell Biol. 2012 Feb;24(1):40-7. doi: 10.1016/j.ceb.2011.12.008. Epub 2012 Jan 3. Curr Opin Cell Biol. 2012. PMID: 22221609 Free PMC article. Review.
-
Measuring Kinetochore-Microtubule Attachment Stability in Cultured Cells.Methods Mol Biol. 2016;1413:147-68. doi: 10.1007/978-1-4939-3542-0_10. Methods Mol Biol. 2016. PMID: 27193848
-
Dynamic regulation of kinetochore-microtubule interaction during mitosis.J Biochem. 2012 Nov;152(5):415-24. doi: 10.1093/jb/mvs109. Epub 2012 Sep 20. J Biochem. 2012. PMID: 22995986 Review.
Cited by
-
Conserved and divergent features of kinetochores and spindle microtubule ends from five species.J Cell Biol. 2013 Feb 18;200(4):459-74. doi: 10.1083/jcb.201209154. J Cell Biol. 2013. PMID: 23420873 Free PMC article.
-
An essential step of kinetochore formation controlled by the SNARE protein Snap29.EMBO J. 2016 Oct 17;35(20):2223-2237. doi: 10.15252/embj.201693991. Epub 2016 Sep 19. EMBO J. 2016. PMID: 27647876 Free PMC article.
-
Cdt1 stabilizes kinetochore-microtubule attachments via an Aurora B kinase-dependent mechanism.J Cell Biol. 2018 Oct 1;217(10):3446-3463. doi: 10.1083/jcb.201705127. Epub 2018 Aug 28. J Cell Biol. 2018. PMID: 30154187 Free PMC article.
-
N-terminus-modified Hec1 suppresses tumour growth by interfering with kinetochore-microtubule dynamics.Oncogene. 2015 Jun;34(25):3325-35. doi: 10.1038/onc.2014.265. Epub 2014 Aug 18. Oncogene. 2015. PMID: 25132262
-
Loss of Centromere Cohesion in Aneuploid Human Oocytes Correlates with Decreased Kinetochore Localization of the Sac Proteins Bub1 and Bubr1.Sci Rep. 2017 Mar 13;7:44001. doi: 10.1038/srep44001. Sci Rep. 2017. PMID: 28287092 Free PMC article.
References
-
- Wendell KL, Wilson L, Jordan MA. Mitotic block in HeLa cells by vinblastine: ultrastructural changes in kinetochore-microtubule attachment and in centrosomes. J Cell Sci. 1993;104(Pt 2):261–274. - PubMed
-
- Cheeseman IM, Desai A. Molecular architecture of the kinetochore–microtubule interface. Nat Rev Mol Cell Biol. 2008;9:33–46. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

