Structural position correlation analysis (SPCA) for protein family
- PMID: 22163002
- PMCID: PMC3230615
- DOI: 10.1371/journal.pone.0028206
Structural position correlation analysis (SPCA) for protein family
Abstract
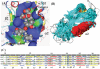
Background: The proteins in a family, which perform the similar biological functions, may have very different amino acid composition, but they must share the similar 3D structures, and keep a stable central region. In the conservative structure region similar biological functions are performed by two or three catalytic residues with the collaboration of several functional residues at key positions. Communication signals are conducted in a position network, adjusting the biological functions in the protein family.
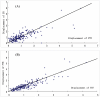
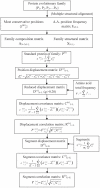
Methodology: A computational approach, namely structural position correlation analysis (SPCA), is developed to analyze the correlation relationship between structural segments (or positions). The basic hypothesis of SPCA is that in a protein family the structural conservation is more important than the sequence conservation, and the local structural changes may contain information of biology functional evolution. A standard protein P(0) is defined in a protein family, which consists of the most-frequent amino acids and takes the average structure of the protein family. The foundational variables of SPCA is the structural position displacements between the standard protein P(0) and individual proteins P(i) of the family. The structural positions are organized as segments, which are the stable units in structural displacements of the protein family. The biological function differences of protein members are determined by the position structural displacements of individual protein P(i) to the standard protein P(0). Correlation analysis is used to analyze the communication network among segments.
Conclusions: The structural position correlation analysis (SPCA) is able to find the correlation relationship among the structural segments (or positions) in a protein family, which cannot be detected by the amino acid sequence and frequency-based methods. The functional communication network among the structural segments (or positions) in protein family, revealed by SPCA approach, well illustrate the distantly allosteric interactions, and contains valuable information for protein engineering study.
Conflict of interest statement
Figures


Similar articles
-
Correlation analysis for protein evolutionary family based on amino acid position mutations and application in PDZ domain.PLoS One. 2010 Oct 6;5(10):e13207. doi: 10.1371/journal.pone.0013207. PLoS One. 2010. PMID: 20949088 Free PMC article.
-
Sequence and structural analysis of cellular retinoic acid-binding proteins reveals a network of conserved hydrophobic interactions.Proteins. 2004 Feb 1;54(2):179-94. doi: 10.1002/prot.10520. Proteins. 2004. PMID: 14696180
-
The ConSurf-HSSP database: the mapping of evolutionary conservation among homologs onto PDB structures.Proteins. 2005 Feb 15;58(3):610-7. doi: 10.1002/prot.20305. Proteins. 2005. PMID: 15614759
-
A tour of structural genomics.Nat Rev Genet. 2001 Oct;2(10):801-9. doi: 10.1038/35093574. Nat Rev Genet. 2001. PMID: 11584296 Review.
-
The construction of an amino acid network for understanding protein structure and function.Amino Acids. 2014 Jun;46(6):1419-39. doi: 10.1007/s00726-014-1710-6. Epub 2014 Mar 13. Amino Acids. 2014. PMID: 24623120 Review.
References
-
- Lichtarge O, Bourne HR, Cohen FE. An evolutionary trace method defines binding surfaces common to protein families. J Mol Biol. 1996;257:342–358. - PubMed
-
- Lockless SW, Ranganathan R. Evolutionarily conserved pathways of energetic connectivity in protein families. Science. 1999;286:295–299. - PubMed
-
- Cover TM, Thomas JA. Elements of information theory. Wiley-Interscience, New York; 2006.
-
- Socolich M, Lockless SW, Russ WP, Lee H, Gardner KH, et al. Evolutionary information for specifying a protein fold. Nature. 2005;437:512–518. - PubMed
-
- Kaczanowski S, Zielenkiewicz P. Why similar protein sequences encode similar three-dimensional structures?. Theor Chem Acc. 2010;125:643–650.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

