Sequence features of E. coli mRNAs affect their degradation
- PMID: 22163312
- PMCID: PMC3233582
- DOI: 10.1371/journal.pone.0028544
Sequence features of E. coli mRNAs affect their degradation
Abstract
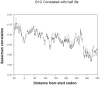
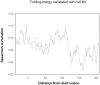
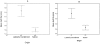
Degradation of mRNA in bacteria is a regulatory mechanism, providing an efficient way to fine-tune protein abundance in response to environmental changes. While the mechanisms responsible for initiation and subsequent propagation of mRNA degradation are well studied, the mRNA features that affect its stability are yet to be elucidated. We calculated three properties for each mRNA in the E. coli transcriptome: G+C content, tRNA adaptation index (tAI) and folding energy. Each of these properties were then correlated with the experimental transcript half life measured for each transcript and detected significant correlations. A sliding window analysis identified the regions that displayed the maximal signal. The correlation between transcript half life and both G+C content and folding energy was strongest at the 5' termini of the mRNAs. Partial correlations showed that each of the parameters contributes separately to mRNA half life. Notably, mRNAs of recently-acquired genes in the E. coli genome, which have a distinct nucleotide composition, tend to be highly stable. This high stability may aid the evolutionary fixation of horizontally acquired genes.
Conflict of interest statement
Figures


References
-
- Coburn GA, Mackie GA. Degradation of mRNA in Escherichia coli: an old problem with some new twists. Prog Nucleic Acid Res Mol Biol. 1999;62:55–108. - PubMed
-
- Shenhar Y, Rasouly A, Biran D, Ron EZ. Adaptation of Escherichi coli to elevated temperatures involves a change in stability of heat shock gene transcripts. Environ Microbiol. 2009;11:2989–2997. - PubMed
-
- Nierlich DP, Murakawa GJ. The decay of bacterial messenger RNA. Prog Nucleic Acid Res Mol Biol. 1996;52:153–216. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

