A novel cold-active β-D-galactosidase from the Paracoccus sp. 32d--gene cloning, purification and characterization
- PMID: 22166118
- PMCID: PMC3268748
- DOI: 10.1186/1475-2859-10-108
A novel cold-active β-D-galactosidase from the Paracoccus sp. 32d--gene cloning, purification and characterization
Abstract
Background: β-D-Galactosidases (EC 3.2.1.23) catalyze the hydrolysis of terminal non-reducing β-D-galactose residues in β-D-galactosides. Cold-active β-D-galactosidases have recently become a focus of attention of researchers and dairy product manufactures owing to theirs ability to: (i) eliminate of lactose from refrigerated milk for people afflicted with lactose intolerance, (ii) convert lactose to glucose and galactose which increase the sweetness of milk and decreases its hydroscopicity, and (iii) eliminate lactose from dairy industry pollutants associated with environmental problems. Moreover, in contrast to commercially available mesophilic β-D-galactosidase from Kluyveromyces lactis the cold-active counterparts could make it possible both to reduce the risk of mesophiles contamination and save energy during the industrial process connected with lactose hydrolysis.
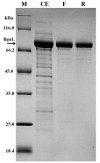
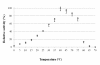
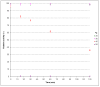
Results: A genomic DNA library was constructed from soil bacterium Paracoccus sp. 32d. Through screening of the genomic DNA library on LB agar plates supplemented with X-Gal, a novel gene encoding a cold-active β-D-galactosidase was isolated. The in silico analysis of the enzyme amino acid sequence revealed that the β-D-galactosidase Paracoccus sp. 32d is a novel member of Glycoside Hydrolase Family 2. However, owing to the lack of a BGal_small_N domain, the domain characteristic for the LacZ enzymes of the GH2 family, it was decided to call the enzyme under study 'BgaL'. The bgaL gene was cloned and expressed in Escherichia coli using the pBAD Expression System. The purified recombinant BgaL consists of two identical subunits with a combined molecular weight of about 160 kDa. The BgaL was optimally active at 40°C and pH 7.5. Moreover, BgaL was able to hydrolyze both lactose and o-nitrophenyl-β-D-galactopyranoside at 10°C with Km values of 2.94 and 1.17 mM and kcat values 43.23 and 71.81 s-1, respectively. One U of the recombinant BgaL would thus be capable hydrolyzing about 97% of the lactose in 1 ml of milk in 24 h at 10°C.
Conclusions: A novel bgaL gene was isolated from Paracoccus sp. 32d encoded a novel cold-active β-D-galactosidase. An E. coli expression system has enabled efficient production of soluble form of BgaL Paracoccus sp. 32d. The amino acid sequence analysis of the BgaL enzyme revealed notable differences in comparison to the result of the amino acid sequences analysis of well-characterized cold-active β-D-galactosidases belonging to Glycoside Hydrolase Family 2. Finally, the enzymatic properties of Paracoccus sp. 32d β-D-galactosidase shows its potential for being applied to development of a new industrial biocatalyst for efficient lactose hydrolysis in milk.
Figures


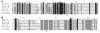


Similar articles
-
Heterologous Production, Purification and Characterization of Two Cold-Active β-d-Galactosidases with Transglycosylation Activity from the Psychrotolerant Arctic Bacterium Arthrobacter sp. S3* Isolated from Spitsbergen Island Soil.Int J Mol Sci. 2024 Dec 12;25(24):13354. doi: 10.3390/ijms252413354. Int J Mol Sci. 2024. PMID: 39769117 Free PMC article.
-
Structural studies of a cold-adapted dimeric β-D-galactosidase from Paracoccus sp. 32d.Acta Crystallogr D Struct Biol. 2016 Sep;72(Pt 9):1049-61. doi: 10.1107/S2059798316012535. Epub 2016 Aug 31. Acta Crystallogr D Struct Biol. 2016. PMID: 27599737
-
Cloning, Expression and Characterization of a Novel Cold-adapted β-galactosidase from the Deep-sea Bacterium Alteromonas sp. ML52.Mar Drugs. 2018 Nov 27;16(12):469. doi: 10.3390/md16120469. Mar Drugs. 2018. PMID: 30486362 Free PMC article.
-
Molecular cloning of lactose genes in dairy lactic streptococci: the phospho-beta-galactosidase and beta-galactosidase genes and their expression products.Biochimie. 1988 Apr;70(4):461-73. doi: 10.1016/0300-9084(88)90083-1. Biochimie. 1988. PMID: 3139067 Review.
-
β-Galactosidase: From its source and applications to its recombinant form.Biotechnol Appl Biochem. 2022 Apr;69(2):612-628. doi: 10.1002/bab.2137. Epub 2021 Mar 16. Biotechnol Appl Biochem. 2022. PMID: 33656174 Review.
Cited by
-
A Review on Psychrophilic β-D-Galactosidases and Their Potential Applications.Appl Biochem Biotechnol. 2023 Apr;195(4):2743-2766. doi: 10.1007/s12010-022-04215-w. Epub 2022 Nov 23. Appl Biochem Biotechnol. 2023. PMID: 36422804 Review.
-
Characterization and Application of a New β-Galactosidase Gal42 From Marine Bacterium Bacillus sp. BY02.Front Microbiol. 2021 Oct 25;12:742300. doi: 10.3389/fmicb.2021.742300. eCollection 2021. Front Microbiol. 2021. PMID: 34759900 Free PMC article.
-
Biotechnological potential of psychrophilic microorganisms as the source of cold-active enzymes in food processing applications.3 Biotech. 2021 Nov;11(11):479. doi: 10.1007/s13205-021-03008-y. Epub 2021 Oct 30. 3 Biotech. 2021. PMID: 34790503 Free PMC article. Review.
-
Characterization of two β-galactosidases LacZ and WspA1 from Nostoc flagelliforme with focus on the latter's central active region.Sci Rep. 2021 Sep 16;11(1):18448. doi: 10.1038/s41598-021-97929-6. Sci Rep. 2021. PMID: 34531460 Free PMC article.
-
Heterologous Production, Purification and Characterization of Two Cold-Active β-d-Galactosidases with Transglycosylation Activity from the Psychrotolerant Arctic Bacterium Arthrobacter sp. S3* Isolated from Spitsbergen Island Soil.Int J Mol Sci. 2024 Dec 12;25(24):13354. doi: 10.3390/ijms252413354. Int J Mol Sci. 2024. PMID: 39769117 Free PMC article.
References
-
- Karasová-Lipovová P, Strnad H, Spiwok V, Malá S, Králová B, Russell NJ. The cloning, purification and characterisation of a cold-active b-galactosidase from the psychrotolerant Antarctic bacterium Arthrobacter sp. C2-2. Enzyme Microb Technol. 2003;33:836–844. doi: 10.1016/S0141-0229(03)00211-4. - DOI
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

