Novel coenzyme B12-dependent interconversion of isovaleryl-CoA and pivalyl-CoA
- PMID: 22167181
- PMCID: PMC3281689
- DOI: 10.1074/jbc.M111.320051
Novel coenzyme B12-dependent interconversion of isovaleryl-CoA and pivalyl-CoA
Abstract
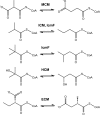
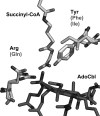
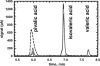
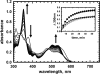
5'-Deoxyadenosylcobalamin (AdoCbl)-dependent isomerases catalyze carbon skeleton rearrangements using radical chemistry. We have recently characterized a fusion protein that comprises the two subunits of the AdoCbl-dependent isobutyryl-CoA mutase flanking a G-protein chaperone and named it isobutyryl-CoA mutase fused (IcmF). IcmF catalyzes the interconversion of isobutyryl-CoA and n-butyryl-CoA, whereas GTPase activity is associated with its G-protein domain. In this study, we report a novel activity associated with IcmF, i.e. the interconversion of isovaleryl-CoA and pivalyl-CoA. Kinetic characterization of IcmF yielded the following values: a K(m) for isovaleryl-CoA of 62 ± 8 μM and V(max) of 0.021 ± 0.004 μmol min(-1) mg(-1) at 37 °C. Biochemical experiments show that an IcmF in which the base specificity loop motif NKXD is modified to NKXE catalyzes the hydrolysis of both GTP and ATP. IcmF is susceptible to rapid inactivation during turnover, and GTP conferred modest protection during utilization of isovaleryl-CoA as substrate. Interestingly, there was no protection from inactivation when either isobutyryl-CoA or n-butyryl-CoA was used as substrate. Detailed kinetic analysis indicated that inactivation is associated with loss of the 5'-deoxyadenosine moiety from the active site, precluding reformation of AdoCbl at the end of the turnover cycle. Under aerobic conditions, oxidation of the cob(II)alamin radical in the inactive enzyme results in accumulation of aquacobalamin. Because pivalic acid found in sludge can be used as a carbon source by some bacteria and isovaleryl-CoA is an intermediate in leucine catabolism, our discovery of a new isomerase activity associated with IcmF expands its metabolic potential.
Figures




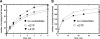
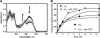
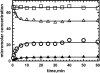
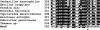
Similar articles
-
Cofactor Editing by the G-protein Metallochaperone Domain Regulates the Radical B12 Enzyme IcmF.J Biol Chem. 2017 Mar 10;292(10):3977-3987. doi: 10.1074/jbc.M117.775957. Epub 2017 Jan 27. J Biol Chem. 2017. PMID: 28130442 Free PMC article.
-
Engineered and Native Coenzyme B12-dependent Isovaleryl-CoA/Pivalyl-CoA Mutase.J Biol Chem. 2015 Aug 14;290(33):20466-76. doi: 10.1074/jbc.M115.646299. Epub 2015 Jul 1. J Biol Chem. 2015. PMID: 26134562 Free PMC article.
-
IcmF is a fusion between the radical B12 enzyme isobutyryl-CoA mutase and its G-protein chaperone.J Biol Chem. 2010 Jan 1;285(1):655-66. doi: 10.1074/jbc.M109.062182. Epub 2009 Oct 28. J Biol Chem. 2010. PMID: 19864421 Free PMC article.
-
Novel B(12)-dependent acyl-CoA mutases and their biotechnological potential.Biochemistry. 2012 Aug 7;51(31):6039-46. doi: 10.1021/bi300827v. Epub 2012 Jul 23. Biochemistry. 2012. PMID: 22803641 Free PMC article. Review.
-
Multiple roles of ATP:cob(I)alamin adenosyltransferases in the conversion of B12 to coenzyme B12.Appl Microbiol Biotechnol. 2010 Sep;88(1):41-8. doi: 10.1007/s00253-010-2773-2. Epub 2010 Jul 31. Appl Microbiol Biotechnol. 2010. PMID: 20677021 Free PMC article. Review.
Cited by
-
Cofactor Editing by the G-protein Metallochaperone Domain Regulates the Radical B12 Enzyme IcmF.J Biol Chem. 2017 Mar 10;292(10):3977-3987. doi: 10.1074/jbc.M117.775957. Epub 2017 Jan 27. J Biol Chem. 2017. PMID: 28130442 Free PMC article.
-
Engineered and Native Coenzyme B12-dependent Isovaleryl-CoA/Pivalyl-CoA Mutase.J Biol Chem. 2015 Aug 14;290(33):20466-76. doi: 10.1074/jbc.M115.646299. Epub 2015 Jul 1. J Biol Chem. 2015. PMID: 26134562 Free PMC article.
-
Allosteric Regulation of Oligomerization by a B12 Trafficking G-Protein Is Corrupted in Methylmalonic Aciduria.Cell Chem Biol. 2019 Jul 18;26(7):960-969.e4. doi: 10.1016/j.chembiol.2019.03.014. Epub 2019 May 2. Cell Chem Biol. 2019. PMID: 31056463 Free PMC article.
-
Phylogenetic analysis of vitamin B12-related metabolism in Mycobacterium tuberculosis.Front Mol Biosci. 2015 Mar 4;2:6. doi: 10.3389/fmolb.2015.00006. eCollection 2015. Front Mol Biosci. 2015. PMID: 25988174 Free PMC article. Review.
-
Role of vitamin B12 on methylmalonyl-CoA mutase activity.J Zhejiang Univ Sci B. 2012 Jun;13(6):423-37. doi: 10.1631/jzus.B1100329. J Zhejiang Univ Sci B. 2012. PMID: 22661206 Free PMC article. Review.
References
-
- Banerjee R. (2003) Radical carbon skeleton rearrangements: catalysis by coenzyme B12-dependent mutases. Chem. Rev. 103, 2083–2094 - PubMed
-
- Ratnatilleke A., Vrijbloed J. W., Robinson J. A. (1999) Cloning and sequencing of the coenzyme B12-binding domain of isobutyryl-CoA mutase from Streptomyces cinnamonensis, reconstitution of mutase activity, and characterization of the recombinant enzyme produced in Escherichia coli. J. Biol. Chem. 274, 31679–31685 - PubMed
-
- Vrijbloed J. W., Zerbe-Burkhardt K., Ratnatilleke A., Grubelnik-Leiser A., Robinson J. A. (1999) Insertional inactivation of methylmalonyl coenzyme A (CoA) mutase and isobutyryl-CoA mutase genes in Streptomyces cinnamonensis: influence on polyketide antibiotic biosynthesis. J. Bacteriol. 181, 5600–5605 - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

