Advancing analytical algorithms and pipelines for billions of microbial sequences
- PMID: 22172529
- PMCID: PMC3273654
- DOI: 10.1016/j.copbio.2011.11.028
Advancing analytical algorithms and pipelines for billions of microbial sequences
Abstract
The vast number of microbial sequences resulting from sequencing efforts using new technologies require us to re-assess currently available analysis methodologies and tools. Here we describe trends in the development and distribution of software for analyzing microbial sequence data. We then focus on one widely used set of methods, dimensionality reduction techniques, which allow users to summarize and compare these vast datasets. We conclude by emphasizing the utility of formal software engineering methods for the development of computational biology tools, and the need for new algorithms for comparing microbial communities. Such large-scale comparisons will allow us to fulfill the dream of rapid integration and comparison of microbial sequence data sets, in a replicable analytical environment, in order to describe the microbial world we inhabit.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures
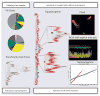
References
-
- Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci U S A. 2011 Mar 15;108(Suppl 1):4516–22. Demonstrates that the Illumina platform's short reads can be used successfully for 16S rRNA profiling in a wide range of environments. - PMC - PubMed
-
- Bartram AK, Lynch MD, Stearns JC, Moreno-Hagelsieb G, Neufeld JD. Generation of multimillion-sequence 16S rRNA gene libraries from complex microbial communities by assembling paired-end illumina reads. Appl Environ Microbiol. 2011 Jun;77(11):3846–52. Demonstrates the advances in sampling depth and quality by using Illumina technologies; also shows the possibility of reducing erroneous sequences by using paired-end reads. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

