Expression of nitrous oxide reductase in Paracoccus denitrificans is regulated by oxygen and nitric oxide through FnrP and NNR
- PMID: 22174385
- PMCID: PMC3541799
- DOI: 10.1099/mic.0.054148-0
Expression of nitrous oxide reductase in Paracoccus denitrificans is regulated by oxygen and nitric oxide through FnrP and NNR
Abstract
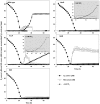
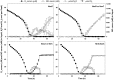
The reductases performing the four steps of denitrification are controlled by a network of transcriptional regulators and ancillary factors responding to intra- and extracellular signals, amongst which are oxygen and N oxides (NO and NO2(-)). Although many components of the regulatory network have been identified, there are gaps in our understanding of their role(s) in controlling the expression of the various reductases, in particular the environmentally important N(2)O reductase (N(2)OR). We investigated denitrification phenotypes of Paracoccus denitrificans mutants deficient in: (i) regulatory proteins (three FNR-type transcriptional regulators, NarR, NNR and FnrP, and NirI, which is involved in transcription activation of the structural nir cluster); (ii) functional enzymes (NO reductase and N(2)OR); or (iii) ancillary factors involved in N(2)O reduction (NirX and NosX). A robotized incubation system allowed us to closely monitor changes in concentrations of oxygen and all gaseous products during the transition from oxic to anoxic respiration. Strains deficient in NO reductase were able to grow during denitrification, despite reaching micromolar concentrations of NO, but were unable to return to oxic respiration. The FnrP mutant showed linear anoxic growth in a medium with nitrate as the sole NO(x), but exponential growth was restored by replacing nitrate with nitrite. We interpret this as nitrite limitation, suggesting dual transcriptional control of respiratory nitrate reductase (NAR) by FnrP and NarR. Mutations in either NirX or NosX did not affect the phenotype, but the double mutant lacked the potential to reduce N(2)O. Finally, we found that FnrP and NNR are alternative and equally effective inducers of N(2)OR.
Figures
References
-
- Bergaust L., Mao Y., Bakken L. R., Frostegård A. (2010). Denitrification response patterns during the transition to anoxic respiration and posttranscriptional effects of suboptimal pH on nitrous [corrected] oxide reductase in Paracoccus denitrificans. Appl Environ Microbiol 76, 6387–6396. 10.1128/AEM.00608-10 - DOI - PMC - PubMed
-
- Bouchal P., Struhárová I., Budinská E., Sedo O., Vyhlídalová T., Zdráhal Z., van Spanning R. J. M., Kucera I. (2010). Unraveling an FNR based regulatory circuit in Paracoccus denitrificans using a proteomics-based approach. Biochim Biophys Acta 1804, 1350–1358. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

