Structural insights into the inhibition of cytosolic 5'-nucleotidase II (cN-II) by ribonucleoside 5'-monophosphate analogues
- PMID: 22174667
- PMCID: PMC3234209
- DOI: 10.1371/journal.pcbi.1002295
Structural insights into the inhibition of cytosolic 5'-nucleotidase II (cN-II) by ribonucleoside 5'-monophosphate analogues
Abstract
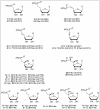
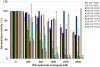
Cytosolic 5'-nucleotidase II (cN-II) regulates the intracellular nucleotide pools within the cell by catalyzing the dephosphorylation of 6-hydroxypurine nucleoside 5'-monophosphates. Beside this physiological function, high level of cN-II expression is correlated with abnormal patient outcome when treated with cytotoxic nucleoside analogues. To identify its specific role in the resistance phenomenon observed during cancer therapy, we screened a particular class of chemical compounds, namely ribonucleoside phosphonates to predict them as potential cN-II inhibitors. These compounds incorporate a chemically and enzymatically stable phosphorus-carbon linkage instead of a regular phosphoester bond. Amongst them, six compounds were predicted as better ligands than the natural substrate of cN-II, inosine 5'-monophosphate (IMP). The study of purine and pyrimidine containing analogues and the introduction of chemical modifications within the phosphonate chain has allowed us to define general rules governing the theoretical affinity of such ligands. The binding strength of these compounds was scrutinized in silico and explained by an impressive number of van der Waals contacts, highlighting the decisive role of three cN-II residues that are Phe 157, His 209 and Tyr 210. Docking predictions were confirmed by experimental measurements of the nucleotidase activity in the presence of the three best available phosphonate analogues. These compounds were shown to induce a total inhibition of the cN-II activity at 2 mM. Altogether, this study emphasizes the importance of the non-hydrolysable phosphonate bond in the design of new competitive cN-II inhibitors and the crucial hydrophobic stacking promoted by three protein residues.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
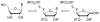




References
-
- Reis MJ. Nucleotidase and its relation to the deamination of nucleotides in the heart and the muscles. Bull Soc Chim Biol. 1934;16:385–399.
-
- Hunsucker SA, Mitchell BS, Spychala J. The 5′-nucleotidases as regulators of nucleotide and drug metabolism. Pharmacol Ther. 2005;107:1–30. - PubMed
-
- Bianchi V, Spychala J. Mammalian 5′-nucleotidases. J Biol Chem. 2003;278:46195–46198. - PubMed
-
- Bretonnet AS, Jordheim LP, Dumontet C, Lancelin JM. Regulation and activity of cytosolic 5′-nucleotidase II. A bifunctional allosteric enzyme of the Haloacid Dehalogenase superfamily involved in cellular metabolism. FEBS Lett. 2005;579:3363–3368. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

