A novel conserved isoform of the ubiquitin ligase UFD2a/UBE4B is expressed exclusively in mature striated muscle cells
- PMID: 22174917
- PMCID: PMC3235170
- DOI: 10.1371/journal.pone.0028861
A novel conserved isoform of the ubiquitin ligase UFD2a/UBE4B is expressed exclusively in mature striated muscle cells
Abstract
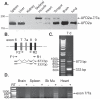
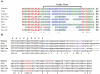
Yeast Ufd2p was the first identified E4 multiubiquitin chain assembly factor. Its vertebrate homologues later referred to as UFD2a, UBE4B or E4B were also shown to have E3 ubiquitin ligase activity. UFD2a function in the brain has been well established in vivo, and in vitro studies have shown that its activity is essential for proper condensation and segregation of chromosomes during mitosis. Here we show that 2 alternative splice forms of UFD2a, UFD2a-7 and -7/7a, are expressed sequentially during myoblast differentiation of C2C12 cell cultures and during cardiotoxin-induced regeneration of skeletal muscle in mice. UFD2a-7 contains an alternate exon 7, and UFD2a-7/7a, the larger of the 2 isoforms, contains an additional novel exon 7a. Analysis of protein or mRNA expression in mice and zebrafish revealed that a similar pattern of isoform switching occurs during developmental myogenesis of cardiac and skeletal muscle. In vertebrates (humans, rodents, zebrafish), UFD2a-7/7a is expressed only in mature striated muscle. This unique tissue specificity is further validated by the conserved presence of 2 muscle-specific splicing regulatory motifs located in the 3' introns of exons 7 and 7a. UFD2a interacts with VCP/p97, an AAA-type ATPase implicated in processes whose functions appear to be regulated, in part, through their interaction with one or more of 15 previously identified cofactors. UFD2a-7/7a did not interact with VCP/p97 in yeast 2-hybrid experiments, which may allow the ATPase to bind cofactors that facilitate its muscle-specific functions. We conclude that the regulated expression of these UFD2a isoforms most likely imparts divergent functions that are important for myogenisis.
Conflict of interest statement
Figures





Similar articles
-
Characterization of the mouse gene for the U-box-type ubiquitin ligase UFD2a.Biochem Biophys Res Commun. 2003 Jan 10;300(2):297-304. doi: 10.1016/s0006-291x(02)02834-6. Biochem Biophys Res Commun. 2003. PMID: 12504083
-
Evolutionary divergence of valosin-containing protein/cell division cycle protein 48 binding interactions among endoplasmic reticulum-associated degradation proteins.FEBS J. 2009 Mar;276(5):1208-20. doi: 10.1111/j.1742-4658.2008.06858.x. FEBS J. 2009. PMID: 19175675
-
The structural and functional basis of the p97/valosin-containing protein (VCP)-interacting motif (VIM): mutually exclusive binding of cofactors to the N-terminal domain of p97.J Biol Chem. 2011 Nov 4;286(44):38679-38690. doi: 10.1074/jbc.M111.274506. Epub 2011 Sep 13. J Biol Chem. 2011. PMID: 21914798 Free PMC article.
-
Insights into adaptor binding to the AAA protein p97.Biochem Soc Trans. 2008 Feb;36(Pt 1):62-7. doi: 10.1042/BST0360062. Biochem Soc Trans. 2008. PMID: 18208387 Review.
-
Valosin-Containing Protein (VCP)/p97: A Prognostic Biomarker and Therapeutic Target in Cancer.Int J Mol Sci. 2021 Sep 21;22(18):10177. doi: 10.3390/ijms221810177. Int J Mol Sci. 2021. PMID: 34576340 Free PMC article. Review.
Cited by
-
PTBP3 modulates P53 expression and promotes colorectal cancer cell proliferation by maintaining UBE4A mRNA stability.Cell Death Dis. 2022 Feb 8;13(2):128. doi: 10.1038/s41419-022-04564-8. Cell Death Dis. 2022. PMID: 35136024 Free PMC article.
-
The E3/E4 ubiquitin conjugation factor UBE4B interacts with and ubiquitinates the HTLV-1 Tax oncoprotein to promote NF-κB activation.PLoS Pathog. 2020 Dec 23;16(12):e1008504. doi: 10.1371/journal.ppat.1008504. eCollection 2020 Dec. PLoS Pathog. 2020. PMID: 33362245 Free PMC article.
-
Skeletal Muscle Regeneration in Cardiotoxin-Induced Muscle Injury Models.Int J Mol Sci. 2022 Nov 2;23(21):13380. doi: 10.3390/ijms232113380. Int J Mol Sci. 2022. PMID: 36362166 Free PMC article. Review.
-
Differential Degradation of TRA2A and PYCR2 Mediated by Ubiquitin E3 Ligase E4B.Front Cell Dev Biol. 2022 May 20;10:833396. doi: 10.3389/fcell.2022.833396. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35669517 Free PMC article.
-
Chaperoning myosin assembly in muscle formation and aging.Worm. 2013 Jul 1;2(3):e25644. doi: 10.4161/worm.25644. Epub 2013 Jul 17. Worm. 2013. PMID: 24778937 Free PMC article.
References
-
- Johnson JM, Castle J, Garrett-Engele P, Kan Z, Loerch PM, et al. Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science. 2003;302:2141–2144. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

