Computational identification and experimental characterization of substrate binding determinants of nucleotide pyrophosphatase/phosphodiesterase 7
- PMID: 22177013
- PMCID: PMC3282672
- DOI: 10.1186/1471-2091-12-65
Computational identification and experimental characterization of substrate binding determinants of nucleotide pyrophosphatase/phosphodiesterase 7
Abstract
Background: Nucleotide pyrophosphatase/phosphodiesterase 7 (NPP7) is the only member of the mammalian NPP enzyme family that has been confirmed to act as a sphingomyelinase, hydrolyzing sphingomyelin (SM) to form phosphocholine and ceramide. NPP7 additionally hydrolyzes lysophosphatidylcholine (LPC), a substrate preference shared with the NPP2/autotaxin(ATX) and NPP6 mammalian family members. This study utilizes a synergistic combination of molecular modeling validated by experimental site-directed mutagenesis to explore the molecular basis for the unique ability of NPP7 to hydrolyze SM.
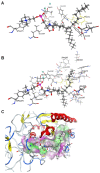
Results: The catalytic function of NPP7 against SM, LPC, platelet activating factor (PAF) and para-nitrophenylphosphorylcholine (pNPPC) is impaired in the F275A mutant relative to wild type NPP7, but different impacts are noted for mutations at other sites. These results are consistent with a previously described role of F275 to interact with the choline headgroup, where all substrates share a common functionality. The L107F mutation showed enhanced hydrolysis of LPC, PAF and pNPPC but reduced hydrolysis of SM. Modeling suggests this difference can be explained by the gain of cation-pi interactions with the choline headgroups of all four substrates, opposed by increased steric crowding against the sphingoid tail of SM. Modeling also revealed that the long and flexible hydrophobic tails of substrates exhibit considerable dynamic flexibility in the binding pocket, reducing the entropic penalty that might otherwise be incurred upon substrate binding.
Conclusions: Substrate recognition by NPP7 includes several important contributions, ranging from cation-pi interactions between F275 and the choline headgroup of all substrates, to tail-group binding pockets that accommodate the inherent flexibility of the lipid hydrophobic tails. Two contributions to the unique ability of NPP7 to hydrolyze SM were identified. First, the second hydrophobic tail of SM occupies a second hydrophobic binding pocket. Second, the leucine residue present at position 107 contrasts with a conserved phenylalanine in NPP enzymes that do not utilize SM as a substrate, consistent with the observed reduction in SM hydrolysis by the NPP7-L107F mutant.
Figures



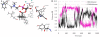

Similar articles
-
Crystal structure of the human alkaline sphingomyelinase provides insights into substrate recognition.J Biol Chem. 2017 Apr 28;292(17):7087-7094. doi: 10.1074/jbc.M116.769273. Epub 2017 Mar 14. J Biol Chem. 2017. PMID: 28292932 Free PMC article.
-
Biochemical and molecular characterization of a novel choline-specific glycerophosphodiester phosphodiesterase belonging to the nucleotide pyrophosphatase/phosphodiesterase family.J Biol Chem. 2005 Jun 17;280(24):23084-93. doi: 10.1074/jbc.M413438200. Epub 2005 Mar 23. J Biol Chem. 2005. PMID: 15788404
-
Understanding the molecular activity of alkaline sphingomyelinase (NPP7) by computer modeling.Biochemistry. 2010 Oct 26;49(42):9096-105. doi: 10.1021/bi101069u. Biochemistry. 2010. PMID: 20839774
-
Alkaline sphingomyelinase (NPP7) in hepatobiliary diseases: A field that needs to be closely studied.World J Hepatol. 2018 Feb 27;10(2):246-253. doi: 10.4254/wjh.v10.i2.246. World J Hepatol. 2018. PMID: 29527260 Free PMC article. Review.
-
Regulation of sphingomyelin metabolism.Pharmacol Rep. 2016 Jun;68(3):570-81. doi: 10.1016/j.pharep.2015.12.008. Epub 2016 Jan 11. Pharmacol Rep. 2016. PMID: 26940196 Review.
Cited by
-
Crystal structure of the human alkaline sphingomyelinase provides insights into substrate recognition.J Biol Chem. 2017 Apr 28;292(17):7087-7094. doi: 10.1074/jbc.M116.769273. Epub 2017 Mar 14. J Biol Chem. 2017. PMID: 28292932 Free PMC article.
References
-
- Duan RD, Bergman T, Xu N, Wu J, Cheng Y, Duan J, Nelander S, Palmberg C, Nilsson A. Identification of human intestinal alkaline sphingomyelinase as a novel ecto-enzyme related to the nucleotide phosphodiesterase family. J Biol Chem. 2003;278(40):38528–38536. doi: 10.1074/jbc.M305437200. - DOI - PubMed
-
- Sakagami H, Aoki J, Natori Y, Nishikawa K, Kakehi Y, Arai H. Biochemical and molecular characterization of a novel choline-specific glycerophosphodiester phosphodiesterase belonging to the nucleotide pyrophosphatase/phosphodiesterase (NPP) family. J Biol Chem. 2005. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

