Cost sensitive hierarchical document classification to triage PubMed abstracts for manual curation
- PMID: 22182279
- PMCID: PMC3314711
- DOI: 10.1186/1471-2105-12-482
Cost sensitive hierarchical document classification to triage PubMed abstracts for manual curation
Abstract
Background: The Immune Epitope Database (IEDB) project manually curates information from published journal articles that describe immune epitopes derived from a wide variety of organisms and associated with different diseases. In the past, abstracts of scientific articles were retrieved by broad keyword queries of PubMed, and were classified as relevant (curatable) or irrelevant (not curatable) to the scope of the database by a Naïve Bayes classifier. The curatable abstracts were subsequently manually classified into categories corresponding to different disease domains. Over the past four years, we have examined how to further improve this approach in order to enhance classification performance and to reduce the need for manual intervention.
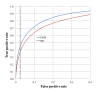
Results: Utilizing 89,884 abstracts classified by a domain expert as curatable or uncuratable, we found that a SVM classifier outperformed the previously used Naïve Bayes classifier for curatability predictions with an AUC of 0.899 and 0.854, respectively. Next, using a non-hierarchical and a hierarchical application of SVM classifiers trained on 22,833 curatable abstracts manually classified into three levels of disease specific categories we demonstrated that a hierarchical application of SVM classifiers outperformed non-hierarchical SVM classifiers for categorization. Finally, to optimize the hierarchical SVM classifiers' error profile for the curation process, cost sensitivity functions were developed to avoid serious misclassifications. We tested our design on a benchmark dataset of 1,388 references and achieved an overall category prediction accuracy of 94.4%, 93.9%, and 82.1% at the three levels of categorization, respectively.
Conclusions: A hierarchical application of SVM algorithms with cost sensitive output weighting enabled high quality reference classification with few serious misclassifications. This enabled us to significantly reduce the manual component of abstract categorization. Our findings are relevant to other databases that are developing their own document classifier schema and the datasets we make available provide large scale real-life benchmark sets for method developers.
Figures

References
-
- Peters B, Sidney J, Bourne P, Bui HH, Doh G, Fleri W, Kronenberg M, Kubo R, Lund O, Nemazee D, Ponomarenko JV, Sathiamurthy M, Schoenberger S, Stewart S, Surko P, Way S, Wilson S, Sette A. The Immune Epitope Database and Analysis Resource: from vision to blueprint. PLoS Biology. 2005;3(3):379–381. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

