A tandem sequence motif acts as a distance-dependent enhancer in a set of genes involved in translation by binding the proteins NonO and SFPQ
- PMID: 22185324
- PMCID: PMC3262029
- DOI: 10.1186/1471-2164-12-624
A tandem sequence motif acts as a distance-dependent enhancer in a set of genes involved in translation by binding the proteins NonO and SFPQ
Abstract
Background: Bioinformatic analyses of expression control sequences in promoters of co-expressed or functionally related genes enable the discovery of common regulatory sequence motifs that might be involved in co-ordinated gene expression. By studying promoter sequences of the human ribosomal protein genes we recently identified a novel highly specific Localized Tandem Sequence Motif (LTSM). In this work we sought to identify additional genes and LTSM-binding proteins to elucidate potential regulatory mechanisms.
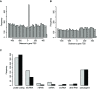
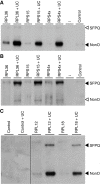
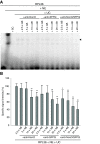
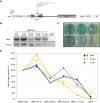
Results: Genome-wide analyses allowed finding a considerable number of additional LTSM-positive genes, the products of which are involved in translation, among them, translation initiation and elongation factors, and 5S rRNA. Electromobility shift assays then showed specific signals demonstrating the binding of protein complexes to LTSM in ribosomal protein gene promoters. Pull-down assays with LTSM-containing oligonucleotides and subsequent mass spectrometric analysis identified the related multifunctional nucleotide binding proteins NonO and SFPQ in the binding complex. Functional characterization then revealed that LTSM enhances the transcriptional activity of the promoters in dependency of the distance from the transcription start site.
Conclusions: Our data demonstrate the power of bioinformatic analyses for the identification of biologically relevant sequence motifs. LTSM and the here found LTSM-binding proteins NonO and SFPQ were discovered through a synergistic combination of bioinformatic and biochemical methods and are regulators of the expression of a set of genes of the translational apparatus in a distance-dependent manner.
Figures

Similar articles
-
SFPQ•NONO and XLF function separately and together to promote DNA double-strand break repair via canonical nonhomologous end joining.Nucleic Acids Res. 2017 Feb 28;45(4):1848-1859. doi: 10.1093/nar/gkw1209. Nucleic Acids Res. 2017. PMID: 27924002 Free PMC article.
-
IGFBP-3 interacts with NONO and SFPQ in PARP-dependent DNA damage repair in triple-negative breast cancer.Cell Mol Life Sci. 2019 May;76(10):2015-2030. doi: 10.1007/s00018-019-03033-4. Epub 2019 Feb 6. Cell Mol Life Sci. 2019. PMID: 30725116 Free PMC article.
-
Involvement of Matrin 3 and SFPQ/NONO in the DNA damage response.Cell Cycle. 2010 Apr 15;9(8):1568-76. doi: 10.4161/cc.9.8.11298. Epub 2010 Apr 15. Cell Cycle. 2010. PMID: 20421735
-
Paraspeckles: nuclear bodies built on long noncoding RNA.J Cell Biol. 2009 Sep 7;186(5):637-44. doi: 10.1083/jcb.200906113. Epub 2009 Aug 31. J Cell Biol. 2009. PMID: 19720872 Free PMC article. Review.
-
Established and Evolving Roles of the Multifunctional Non-POU Domain-Containing Octamer-Binding Protein (NonO) and Splicing Factor Proline- and Glutamine-Rich (SFPQ).J Dev Biol. 2024 Jan 5;12(1):3. doi: 10.3390/jdb12010003. J Dev Biol. 2024. PMID: 38248868 Free PMC article. Review.
Cited by
-
The glycolytic enzyme ALDOA and the exon junction complex protein RBM8A are regulators of ribosomal biogenesis.Front Cell Dev Biol. 2022 Sep 14;10:954358. doi: 10.3389/fcell.2022.954358. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 36187487 Free PMC article.
-
Polymorphisms and genetic effects of PRLR, MOGAT1, MINPP1 and CHUK genes on milk fatty acid traits in Chinese Holstein.BMC Genet. 2019 Aug 16;20(1):69. doi: 10.1186/s12863-019-0769-1. BMC Genet. 2019. PMID: 31419940 Free PMC article.
-
An alternative cytoplasmic SFPQ isoform with reduced phase separation potential is up-regulated in ALS.Sci Adv. 2025 Aug 22;11(34):eadt4814. doi: 10.1126/sciadv.adt4814. Epub 2025 Aug 22. Sci Adv. 2025. PMID: 40845103 Free PMC article.
-
Characterization of bursa subacromialis-derived mesenchymal stem cells.Stem Cell Res Ther. 2015 Jun 3;6(1):114. doi: 10.1186/s13287-015-0104-3. Stem Cell Res Ther. 2015. PMID: 26036250 Free PMC article.
-
Double-strand break repair deficiency in NONO knockout murine embryonic fibroblasts and compensation by spontaneous upregulation of the PSPC1 paralog.Nucleic Acids Res. 2014 Sep;42(15):9771-80. doi: 10.1093/nar/gku650. Epub 2014 Aug 6. Nucleic Acids Res. 2014. PMID: 25100870 Free PMC article.
References
-
- Roepcke S, Zhi D, Vingron M, Arndt PF. Identification of highly specific localized sequence motifs in human ribosomal protein gene promoters. Gene. 2006;365:48–56. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

