Queueing up for enzymatic processing: correlated signaling through coupled degradation
- PMID: 22186735
- PMCID: PMC3737734
- DOI: 10.1038/msb.2011.94
Queueing up for enzymatic processing: correlated signaling through coupled degradation
Abstract
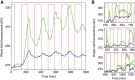
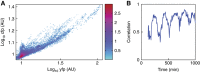
High-throughput technologies have led to the generation of complex wiring diagrams as a post-sequencing paradigm for depicting the interactions between vast and diverse cellular species. While these diagrams are useful for analyzing biological systems on a large scale, a detailed understanding of the molecular mechanisms that underlie the observed network connections is critical for the further development of systems and synthetic biology. Here, we use queueing theory to investigate how 'waiting lines' can lead to correlations between protein 'customers' that are coupled solely through a downstream set of enzymatic 'servers'. Using the E. coli ClpXP degradation machine as a model processing system, we observe significant cross-talk between two networks that are indirectly coupled through a common set of processors. We further illustrate the implications of enzymatic queueing using a synthetic biology application, in which two independent synthetic networks demonstrate synchronized behavior when common ClpXP machinery is overburdened. Our results demonstrate that such post-translational processes can lead to dynamic connections in cellular networks and may provide a mechanistic understanding of existing but currently inexplicable links.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures




References
-
- Bougdour A, Cunning C, Baptiste P, Elliott T, Gottesman S (2008) Multiple pathways for regulation of σS (RpoS) stability in Escherichia coli via the action of multiple anti-adaptors. Mol Microbiol 68: 298–313 - PubMed
-
- Bramson M (1998) State space collapse with application to heavy traffic limits for multiclass queueing networks. Queueing Syst 30: 89–148
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

