CRISPR loci reveal networks of gene exchange in archaea
- PMID: 22188759
- PMCID: PMC3285040
- DOI: 10.1186/1745-6150-6-65
CRISPR loci reveal networks of gene exchange in archaea
Abstract
Background: CRISPR (Clustered, Regularly, Interspaced, Short, Palindromic Repeats) loci provide prokaryotes with an adaptive immunity against viruses and other mobile genetic elements. CRISPR arrays can be transcribed and processed into small crRNA molecules, which are then used by the cell to target the foreign nucleic acid. Since spacers are accumulated by active CRISPR/Cas systems, the sequences of these spacers provide a record of the past "infection history" of the organism.
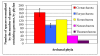
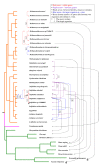
Results: Here we analyzed all currently known spacers present in archaeal genomes and identified their source by DNA similarity. While nearly 50% of archaeal spacers matched mobile genetic elements, such as plasmids or viruses, several others matched chromosomal genes of other organisms, primarily other archaea. Thus, networks of gene exchange between archaeal species were revealed by the spacer analysis, including many cases of inter-genus and inter-species gene transfer events. Spacers that recognize viral sequences tend to be located further away from the leader sequence, implying that there exists a selective pressure for their retention.
Conclusions: CRISPR spacers provide direct evidence for extensive gene exchange in archaea, especially within genera, and support the current dogma where the primary role of the CRISPR/Cas system is anti-viral and anti-plasmid defense.
Open peer review: This article was reviewed by: Profs. W. Ford Doolittle, John van der Oost, Christa Schleper (nominated by board member Prof. J Peter Gogarten).
Figures



References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

