Topoisomerase inhibitors unsilence the dormant allele of Ube3a in neurons
- PMID: 22190039
- PMCID: PMC3257422
- DOI: 10.1038/nature10726
Topoisomerase inhibitors unsilence the dormant allele of Ube3a in neurons
Abstract
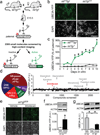
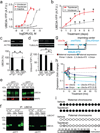
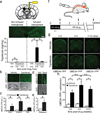
Angelman syndrome is a severe neurodevelopmental disorder caused by deletion or mutation of the maternal allele of the ubiquitin protein ligase E3A (UBE3A). In neurons, the paternal allele of UBE3A is intact but epigenetically silenced, raising the possibility that Angelman syndrome could be treated by activating this silenced allele to restore functional UBE3A protein. Using an unbiased, high-content screen in primary cortical neurons from mice, we identify twelve topoisomerase I inhibitors and four topoisomerase II inhibitors that unsilence the paternal Ube3a allele. These drugs included topotecan, irinotecan, etoposide and dexrazoxane (ICRF-187). At nanomolar concentrations, topotecan upregulated catalytically active UBE3A in neurons from maternal Ube3a-null mice. Topotecan concomitantly downregulated expression of the Ube3a antisense transcript that overlaps the paternal copy of Ube3a. These results indicate that topotecan unsilences Ube3a in cis by reducing transcription of an imprinted antisense RNA. When administered in vivo, topotecan unsilenced the paternal Ube3a allele in several regions of the nervous system, including neurons in the hippocampus, neocortex, striatum, cerebellum and spinal cord. Paternal expression of Ube3a remained elevated in a subset of spinal cord neurons for at least 12 weeks after cessation of topotecan treatment, indicating that transient topoisomerase inhibition can have enduring effects on gene expression. Although potential off-target effects remain to be investigated, our findings suggest a therapeutic strategy for reactivating the functional but dormant allele of Ube3a in patients with Angelman syndrome.
Figures
Comment in
-
Angelman syndrome: Drugs to awaken a paternal gene.Nature. 2011 Dec 21;481(7380):150-2. doi: 10.1038/nature10784. Nature. 2011. PMID: 22190038 Free PMC article.
-
Neurodevelopmental disorders: Unsilencing dormant Ube3a--hope for Angelman syndrome?Nat Rev Neurol. 2012 Jan 24;8(2):62. doi: 10.1038/nrneurol.2012.2. Nat Rev Neurol. 2012. PMID: 22270025 No abstract available.
References
-
- Kishino T, Lalande M, Wagstaff J. UBE3A/E6-AP mutations cause Angelman syndrome. Nature Genetics. 1997;15:70–73. - PubMed
-
- Matsuura T, et al. De novo truncating mutations in E6-AP ubiquitin-protein ligase gene (UBE3A) in Angelman syndrome. Nature Genetics. 1997;15:74–77. - PubMed
-
- Albrecht U, et al. Imprinted expression of the murine Angelman syndrome gene, Ube3a, in hippocampal and Purkinje neurons. Nature Genetics. 1997;17:75–78. - PubMed
-
- Rougeulle C, Glatt H, Lalande M. The Angelman syndrome candidate gene, UBE3A/E6-AP, is imprinted in brain. Nature Genetics. 1997;17:14–15. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 MH093372/MH/NIMH NIH HHS/United States
- R01 NS060725/NS/NINDS NIH HHS/United States
- T32HD040127-07/HD/NICHD NIH HHS/United States
- 5P30NS045892/NS/NINDS NIH HHS/United States
- P30 NS045892/NS/NINDS NIH HHS/United States
- F32 NS067712/NS/NINDS NIH HHS/United States
- R01MH093372/MH/NIMH NIH HHS/United States
- T32 HD040127/HD/NICHD NIH HHS/United States
- 5F32NS067712/NS/NINDS NIH HHS/United States
- R01EY018323/EY/NEI NIH HHS/United States
- R01NS060725/NS/NINDS NIH HHS/United States
- P30 HD003110/HD/NICHD NIH HHS/United States
- R01 EY018323/EY/NEI NIH HHS/United States
- R01NS067688/NS/NINDS NIH HHS/United States
- P30HD03110/HD/NICHD NIH HHS/United States
- HHSN-271-2008-00025-C/PHS HHS/United States
- R01 NS067688/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
