Loss-of-function thrombospondin-1 mutations in familial pulmonary hypertension
- PMID: 22198906
- PMCID: PMC3311532
- DOI: 10.1152/ajplung.00282.2011
Loss-of-function thrombospondin-1 mutations in familial pulmonary hypertension
Abstract
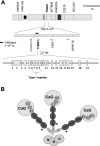
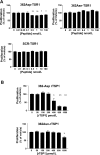
Most patients with familial pulmonary arterial hypertension (FPAH) carry mutations in the bone morphogenic protein receptor 2 gene (BMPR2). Yet carriers have only a 20% risk of disease, suggesting that other factors influence penetrance. Thrombospondin-1 (TSP1) regulates activation of TGF-β and inhibits endothelial and smooth muscle cell proliferation, pathways coincidentally altered in pulmonary arterial hypertension (PAH). To determine whether a subset of FPAH patients also have mutations in the TSP1 gene (THBS1) we resequenced the type I repeats of THBS1 encoding the TGF-β regulation and cell growth inhibition domains in 60 FPAH probands, 70 nonfamilial PAH subjects, and in large control groups. We identified THBS1 mutations in three families: a novel missense mutation in two (Asp362Asn), and an intronic mutation in a third (IVS8+255 G/A). Neither mutation was detected in population controls. Mutant 362Asn TSP1 had less than half of the ability of wild-type TSP1 to activate TGF-β. Mutant 362Asn TSP1 also lost the ability to inhibit growth of pulmonary arterial smooth muscle cells and was over threefold less effective at inhibiting endothelial cell growth. The IVS8+255 G/A mutation decreased and/or eliminated local binding of the transcription factors SP1 and MAZ but did not affect RNA splicing. These novel mutations implicate THBS1 as a modifier gene in FPAH. These THBS1 mutations have implications in the genetic evaluation of FPAH patients. However, since FPAH is rare, these data are most relevant as evidence for the importance of TSP1 in pulmonary vascular homeostasis. Further examination of THBS1 in the pathogenesis of PAH is warranted.
Figures





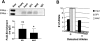
References
-
- Abe M, Harpel JG, Metz CN, Nunes I, Loskutoff DJ, Rifkin DB. An assay for transforming growth factor-beta using cells transfected with a plasminogen activator inhibitor-1 promoter-luciferase construct. Anal Biochem 216: 276–284, 1994 - PubMed
-
- Botella LM, Sanchez-Elsner T, Rius C, Corbi A, Bernabeu C. Identification of a critical Sp1 site within the endoglin promoter and its involvement in the transforming growth factor-beta stimulation. J Biol Chem 276: 34486–34494, 2001 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

