Testing gene-environment interaction in large-scale case-control association studies: possible choices and comparisons
- PMID: 22199027
- PMCID: PMC3286201
- DOI: 10.1093/aje/kwr367
Testing gene-environment interaction in large-scale case-control association studies: possible choices and comparisons
Abstract
Several methods for screening gene-environment interaction have recently been proposed that address the issue of using gene-environment independence in a data-adaptive way. In this report, the authors present a comparative simulation study of power and type I error properties of 3 classes of procedures: 1) the standard 1-step case-control method; 2) the case-only method that requires an assumption of gene-environment independence for the underlying population; and 3) a variety of hybrid methods, including empirical-Bayes, 2-step, and model averaging, that aim at gaining power by exploiting the assumption of gene-environment independence and yet can protect against false positives when the independence assumption is violated. These studies suggest that, although the case-only method generally has maximum power, it has the potential to create substantial false positives in large-scale studies even when a small fraction of markers are associated with the exposure under study in the underlying population. All the hybrid methods perform well in protecting against such false positives and yet can retain substantial power advantages over standard case-control tests. The authors conclude that, for future genome-wide scans for gene-environment interactions, major power gain is possible by using alternatives to standard case-control analysis. Whether a case-only type scan or one of the hybrid methods should be used depends on the strength and direction of gene-environment interaction and association, the level of tolerance for false positives, and the nature of replication strategies.
Figures
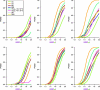
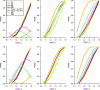
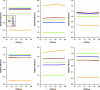
Comment in
-
Invited commentary: GE-Whiz! Ratcheting gene-environment studies up to the whole genome and the whole exposome.Am J Epidemiol. 2012 Feb 1;175(3):203-7; discussion 208-9. doi: 10.1093/aje/kwr365. Epub 2011 Dec 22. Am J Epidemiol. 2012. PMID: 22199029 Free PMC article.

