Unique structural features and sequence motifs of proline utilization A (PutA)
- PMID: 22201760
- PMCID: PMC3715380
- DOI: 10.2741/3943
Unique structural features and sequence motifs of proline utilization A (PutA)
Abstract
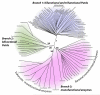
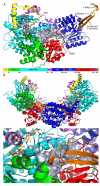
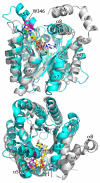
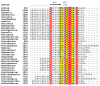
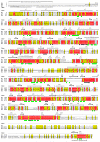
Proline utilization A proteins (PutAs) are bifunctional enzymes that catalyze the oxidation of proline to glutamate using spatially separated proline dehydrogenase and pyrroline-5-carboxylate dehydrogenase active sites. Here we use the crystal structure of the minimalist PutA from Bradyrhizobium japonicum (BjPutA) along with sequence analysis to identify unique structural features of PutAs. This analysis shows that PutAs have secondary structural elements and domains not found in the related monofunctional enzymes. Some of these extra features are predicted to be important for substrate channeling in BjPutA. Multiple sequence alignment analysis shows that some PutAs have a 17-residue conserved motif in the C-terminal 20-30 residues of the polypeptide chain. The BjPutA structure shows that this motif helps seal the internal substrate-channeling cavity from the bulk medium. Finally, it is shown that some PutAs have a 100-200 residue domain of unknown function in the C-terminus that is not found in minimalist PutAs. Remote homology detection suggests that this domain is homologous to the oligomerization beta-hairpin and Rossmann fold domain of BjPutA.
Figures
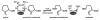


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

