Novel features of ARS selection in budding yeast Lachancea kluyveri
- PMID: 22204614
- PMCID: PMC3306766
- DOI: 10.1186/1471-2164-12-633
Novel features of ARS selection in budding yeast Lachancea kluyveri
Abstract
Background: The characterization of DNA replication origins in yeast has shed much light on the mechanisms of initiation of DNA replication. However, very little is known about the evolution of origins or the evolution of mechanisms through which origins are recognized by the initiation machinery. This lack of understanding is largely due to the vast evolutionary distances between model organisms in which origins have been examined.
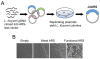
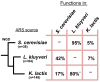
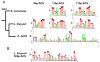
Results: In this study we have isolated and characterized autonomously replicating sequences (ARSs) in Lachancea kluyveri - a pre-whole genome duplication (WGD) budding yeast. Through a combination of experimental work and rigorous computational analysis, we show that L. kluyveri ARSs require a sequence that is similar but much longer than the ARS Consensus Sequence well defined in Saccharomyces cerevisiae. Moreover, compared with S. cerevisiae and K. lactis, the replication licensing machinery in L. kluyveri seems more tolerant to variations in the ARS sequence composition. It is able to initiate replication from almost all S. cerevisiae ARSs tested and most Kluyveromyces lactis ARSs. In contrast, only about half of the L. kluyveri ARSs function in S. cerevisiae and less than 10% function in K. lactis.
Conclusions: Our findings demonstrate a replication initiation system with novel features and underscore the functional diversity within the budding yeasts. Furthermore, we have developed new approaches for analyzing biologically functional DNA sequences with ill-defined motifs.
Figures



Similar articles
-
Isolation and sequence analysis of a K. lactis chromosomal DNA element able to autonomously replicate in S. cerevisiae and K. lactis.Yeast. 1990 Jan-Feb;6(1):69-76. doi: 10.1002/yea.320060108. Yeast. 1990. PMID: 2180237
-
Origin replication complex binding, nucleosome depletion patterns, and a primary sequence motif can predict origins of replication in a genome with epigenetic centromeres.mBio. 2014 Sep 2;5(5):e01703-14. doi: 10.1128/mBio.01703-14. mBio. 2014. PMID: 25182328 Free PMC article.
-
Prediction of Saccharomyces cerevisiae replication origins.Genome Biol. 2004;5(4):R22. doi: 10.1186/gb-2004-5-4-r22. Epub 2004 Mar 4. Genome Biol. 2004. PMID: 15059255 Free PMC article.
-
Structure, replication efficiency and fragility of yeast ARS elements.Res Microbiol. 2012 May;163(4):243-53. doi: 10.1016/j.resmic.2012.03.003. Epub 2012 Mar 28. Res Microbiol. 2012. PMID: 22504206 Review.
-
The dual role of autonomously replicating sequences as origins of replication and as silencers.Curr Genet. 2009 Aug;55(4):357-63. doi: 10.1007/s00294-009-0265-7. Epub 2009 Jul 26. Curr Genet. 2009. PMID: 19633981 Review.
Cited by
-
Conservation of a DNA Replication Motif among Phylogenetically Distant Budding Yeast Species.Genome Biol Evol. 2021 Jul 6;13(7):evab137. doi: 10.1093/gbe/evab137. Genome Biol Evol. 2021. PMID: 34132803 Free PMC article.
-
An autonomously replicating sequence for use in a wide range of budding yeasts.FEMS Yeast Res. 2014 Mar;14(2):364-7. doi: 10.1111/1567-1364.12123. Epub 2013 Dec 2. FEMS Yeast Res. 2014. PMID: 24205893 Free PMC article.
-
Maintaining replication origins in the face of genomic change.Genome Res. 2012 Oct;22(10):1940-52. doi: 10.1101/gr.138248.112. Epub 2012 Jun 4. Genome Res. 2012. PMID: 22665441 Free PMC article.
-
The evolution of the temporal program of genome replication.Nat Commun. 2018 Jun 6;9(1):2199. doi: 10.1038/s41467-018-04628-4. Nat Commun. 2018. PMID: 29875360 Free PMC article.
-
The spatiotemporal program of replication in the genome of Lachancea kluyveri.Genome Biol Evol. 2013;5(2):370-88. doi: 10.1093/gbe/evt014. Genome Biol Evol. 2013. PMID: 23355306 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

