Role of glutamate 64 in the activation of the prodrug 5-fluorocytosine by yeast cytosine deaminase
- PMID: 22208667
- PMCID: PMC3272785
- DOI: 10.1021/bi201540z
Role of glutamate 64 in the activation of the prodrug 5-fluorocytosine by yeast cytosine deaminase
Abstract
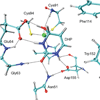
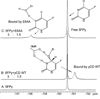
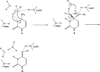
Yeast cytosine deaminase (yCD) catalyzes the hydrolytic deamination of cytosine to uracil as well as the deamination of the prodrug 5-fluorocytosine (5FC) to the anticancer drug 5-fluorouracil. In this study, the role of Glu64 in the activation of the prodrug 5FC was investigated by site-directed mutagenesis, biochemical, nuclear magnetic resonance (NMR), and computational studies. Steady-state kinetics studies showed that the mutation of Glu64 causes a dramatic decrease in k(cat) and a dramatic increase in K(m), indicating Glu64 is important for both binding and catalysis in the activation of 5FC. (19)F NMR experiments showed that binding of the inhibitor 5-fluoro-1H-pyrimidin-2-one (5FPy) to the wild-type yCD causes an upfield shift, indicating that the bound inhibitor is in the hydrated form, mimicking the transition state or the tetrahedral intermediate in the activation of 5FC. However, binding of 5FPy to the E64A mutant enzyme causes a downfield shift, indicating that the bound 5FPy remains in an unhydrated form in the complex with the mutant enzyme. (1)H and (15)N NMR analysis revealed trans-hydrogen bond D/H isotope effects on the hydrogen of the amide of Glu64, indicating that the carboxylate of Glu64 forms two hydrogen bonds with the hydrated 5FPy. ONIOM calculations showed that the wild-type yCD complex with the hydrated form of the inhibitor 1H-pyrimidin-2-one is more stable than the initial binding complex, and in contrast, with the E64A mutant enzyme, the hydrated inhibitor is no longer favored and the conversion has a higher activation energy, as well. The hydrated inhibitor is stabilized in the wild-type yCD by two hydrogen bonds between it and the carboxylate of Glu64 as revealed by (1)H and (15)N NMR analysis. To explore the functional role of Glu64 in catalysis, we investigated the deamination of cytosine catalyzed by the E64A mutant by ONIOM calculations. The results showed that without the assistance of Glu64, both proton transfers before and after the formation of the tetrahedral reaction intermediate become partially rate-limiting steps. The results of the experimental and computational studies together indicate that Glu64 plays a critical role in both the binding and the chemical transformation in the conversion of the prodrug 5FC to the anticancer drug 5-fluorouracil.
Figures




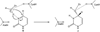



Similar articles
-
Product release is rate-limiting in the activation of the prodrug 5-fluorocytosine by yeast cytosine deaminase.Biochemistry. 2005 Apr 19;44(15):5940-7. doi: 10.1021/bi050095n. Biochemistry. 2005. PMID: 15823054
-
Yeast cytosine deaminase mutants with increased thermostability impart sensitivity to 5-fluorocytosine.J Mol Biol. 2008 Mar 28;377(3):854-69. doi: 10.1016/j.jmb.2008.01.002. Epub 2008 Jan 11. J Mol Biol. 2008. PMID: 18291415 Free PMC article.
-
Product release mechanism and the complete enzyme catalysis cycle in yeast cytosine deaminase (yCD): A computational study.Biochim Biophys Acta Proteins Proteom. 2017 Aug;1865(8):1020-1029. doi: 10.1016/j.bbapap.2017.05.001. Epub 2017 May 3. Biochim Biophys Acta Proteins Proteom. 2017. PMID: 28478051
-
Escherichia coli cytosine deaminase: Structural and biotechnological aspects.Biotechnol Appl Biochem. 2024 Feb;71(1):5-16. doi: 10.1002/bab.2516. Epub 2023 Sep 24. Biotechnol Appl Biochem. 2024. PMID: 37743549 Review.
-
Emerging implications of nonmammalian cytosine deaminases on cancer therapeutics.Appl Biochem Biotechnol. 2012 Aug;167(7):2103-16. doi: 10.1007/s12010-012-9746-0. Epub 2012 Jun 7. Appl Biochem Biotechnol. 2012. PMID: 22673971 Review.
Cited by
-
Asymmetrical dose responses shape the evolutionary trade-off between antifungal resistance and nutrient use.Nat Ecol Evol. 2022 Oct;6(10):1501-1515. doi: 10.1038/s41559-022-01846-4. Epub 2022 Sep 1. Nat Ecol Evol. 2022. PMID: 36050399
-
Compensatory mutations potentiate constructive neutral evolution by gene duplication.bioRxiv [Preprint]. 2024 Feb 13:2024.02.12.579783. doi: 10.1101/2024.02.12.579783. bioRxiv. 2024. Update in: Science. 2024 Aug 16;385(6710):770-775. doi: 10.1126/science.ado5719. PMID: 38405844 Free PMC article. Updated. Preprint.
-
Theoretical insights into the mechanism of redox switch in heat shock protein Hsp33.J Biol Inorg Chem. 2015 Apr;20(3):555-62. doi: 10.1007/s00775-015-1240-z. Epub 2015 Jan 31. J Biol Inorg Chem. 2015. PMID: 25637463
-
Engineering the Active Site Lid Dynamics to Improve the Catalytic Efficiency of Yeast Cytosine Deaminase.Int J Mol Sci. 2023 Apr 1;24(7):6592. doi: 10.3390/ijms24076592. Int J Mol Sci. 2023. PMID: 37047565 Free PMC article.
-
Oxidation reactivity of zinc-cysteine clusters in metallothionein.J Biol Inorg Chem. 2013 Mar;18(3):333-42. doi: 10.1007/s00775-013-0977-5. Epub 2013 Jan 20. J Biol Inorg Chem. 2013. PMID: 23334196
References
-
- Aghi M, Hochberg F, Breakefield XO. Prodrug activation enzymes in cancer gene therapy. J. Gene Med. 2000;2:148–164. - PubMed
-
- Greco O, Dachs GU. Gene directed enzyme/prodrug therapy of cancer: Historical appraisal and future prospectives. J. Cell. Physiol. 2001;187:22–36. - PubMed
-
- Both GW. Recent progress in gene-directed enzyme prodrug therapy: An emerging cancer treatment. Curr. Opin. Mol. Ther. 2009;11:421–432. - PubMed
-
- Ireton GC, McDermott G, Black ME, Stoddard BL. The structure of Escherichia coli cytosine deaminase. J. Mol. Biol. 2002;315:687–697. - PubMed
-
- Ireton GC, Black ME, Stoddard BL. The 1.14 angstrom crystal structure of yeast cytosine deaminase: Evolution of nucleotide salvage enzymes and implications for genetic chemotherapy. Structure. 2003;11:961–972. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

