Transcriptome and functional analysis of mating in the basidiomycete Schizophyllum commune
- PMID: 22210832
- PMCID: PMC3346423
- DOI: 10.1128/EC.05214-11
Transcriptome and functional analysis of mating in the basidiomycete Schizophyllum commune
Abstract
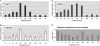
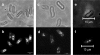
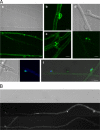
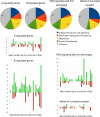
In this study, we undertook a functional characterization and transcriptome analysis that enabled a comprehensive study of the mating type loci of the mushroom Schizophyllum commune. Induced expression of both the bar2 receptor and the bap2(2) pheromone gene within 6 to 12 h after mates' contact was demonstrated by quantitative real-time PCR. Similar temporal expression patterns were confirmed for the allelic bbr1 receptor and bbp1 pheromone-encoding genes by Northern hybridization. Interestingly, the fusion of clamp connections to the subterminal cell was delayed in mating interactions in which one of the compatible partners expressed the bar2 receptor with a truncated C terminus. This developmental delay allowed the visualization of a green fluorescent protein (Gfp)-labeled truncated receptor at the cell periphery, consistent with a localization in the plasma membrane of unfused pseudoclamps. This finding does not support hypotheses envisioning a receptor localization to the nuclear membrane facilitating recognition between the two different nuclei present in each dikaryotic cell. Rather, Gfp fluorescence observed in such pseudoclamps indicated a role of receptor-pheromone interaction in clamp fusion. Transcriptome changes associated with mating interactions were analyzed in order to identify a role for pheromone-receptor interactions. We detected a total of 89 genes that were transcriptionally regulated in a mating type locus A-dependent manner, employing a cutoff of 5-fold changes in transcript abundance. Upregulation in cell cycle-related genes and downregulation of genes involved in metabolism were seen with this set of experiments. In contrast, mating type locus B-dependent transcriptome changes were observed in 208 genes, with a specific impact on genes related to cell wall and membrane metabolism, stress response, and the redox status of the cell.
Figures




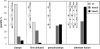

Similar articles
-
Identification and functional analysis of pheromone and receptor genes in the B3 mating locus of Pleurotus eryngii.PLoS One. 2014 Aug 18;9(8):e104693. doi: 10.1371/journal.pone.0104693. eCollection 2014. PLoS One. 2014. PMID: 25133513 Free PMC article.
-
A central role for Ras1 in morphogenesis of the basidiomycete Schizophyllum commune.Eukaryot Cell. 2013 Jun;12(6):941-52. doi: 10.1128/EC.00355-12. Epub 2013 Apr 19. Eukaryot Cell. 2013. PMID: 23606288 Free PMC article.
-
The regulator of G-protein signalling Thn1 links pheromone response to volatile production in Schizophyllum commune.Environ Microbiol. 2018 Oct;20(10):3684-3699. doi: 10.1111/1462-2920.14369. Epub 2018 Sep 17. Environ Microbiol. 2018. PMID: 30062773
-
Mating types and pheromone recognition in the Homobasidiomycete schizophyllum commune.Fungal Genet Biol. 1999 Jul-Aug;27(2-3):146-52. doi: 10.1006/fgbi.1999.1129. Fungal Genet Biol. 1999. PMID: 10441440 Review.
-
Mushrooms by magic: making connections between signal transduction and fruiting body development in the basidiomycete fungus Schizophyllum commune.FEMS Microbiol Lett. 2006 Sep;262(1):1-8. doi: 10.1111/j.1574-6968.2006.00341.x. FEMS Microbiol Lett. 2006. PMID: 16907732 Review.
Cited by
-
Identification and functional analysis of pheromone and receptor genes in the B3 mating locus of Pleurotus eryngii.PLoS One. 2014 Aug 18;9(8):e104693. doi: 10.1371/journal.pone.0104693. eCollection 2014. PLoS One. 2014. PMID: 25133513 Free PMC article.
-
Comparative analysis of the secretomes of Schizophyllum commune and other wood-decay basidiomycetes during solid-state fermentation reveals its unique lignocellulose-degrading enzyme system.Biotechnol Biofuels. 2016 Feb 20;9:42. doi: 10.1186/s13068-016-0461-x. eCollection 2016. Biotechnol Biofuels. 2016. PMID: 26900401 Free PMC article.
-
Sexual selection in hermaphrodites, sperm and broadcast spawners, plants and fungi.Philos Trans R Soc Lond B Biol Sci. 2016 Oct 19;371(1706):20150541. doi: 10.1098/rstb.2015.0541. Philos Trans R Soc Lond B Biol Sci. 2016. PMID: 27619704 Free PMC article. Review.
-
An Overview of the Function and Maintenance of Sexual Reproduction in Dikaryotic Fungi.Front Microbiol. 2018 Mar 21;9:503. doi: 10.3389/fmicb.2018.00503. eCollection 2018. Front Microbiol. 2018. PMID: 29619017 Free PMC article. Review.
-
The role of pheromone receptors for communication and mating in Hypocrea jecorina (Trichoderma reesei).Fungal Genet Biol. 2012 Oct;49(10):814-24. doi: 10.1016/j.fgb.2012.07.004. Epub 2012 Aug 3. Fungal Genet Biol. 2012. PMID: 22884620 Free PMC article.
References
-
- Attwood TK, Findlay JBC. 1994. Fingerprinting G-protein-coupled receptors. Protein Eng. 7:195–203 - PubMed
-
- Ballon DR, et al. 2006. DEP-domain-mediated regulation of GPCR signaling responses. Cell 126:1079–1093 - PubMed
-
- Bölker M, Urban M, Kahmann R. 1992. The A mating type locus of U. maydis specifies cell signaling components. Cell 68:441–450 - PubMed
-
- Debuchy R. 1999. Internuclear recognition: a possible connection between euascomycetes and homobasidiomycetes. Fungal Genet. Biol. 27:218–223 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

