Role of polynucleotide kinase/phosphatase in mitochondrial DNA repair
- PMID: 22210862
- PMCID: PMC3333865
- DOI: 10.1093/nar/gkr1245
Role of polynucleotide kinase/phosphatase in mitochondrial DNA repair
Abstract
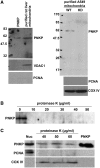
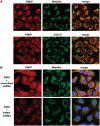
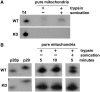
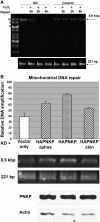
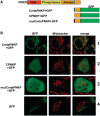
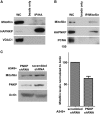
Mutations in mitochondrial DNA (mtDNA) are implicated in a broad range of human diseases and in aging. Compared to nuclear DNA, mtDNA is more highly exposed to oxidative damage due to its proximity to the respiratory chain and the lack of protection afforded by chromatin-associated proteins. While repair of oxidative damage to the bases in mtDNA through the base excision repair pathway has been well studied, the repair of oxidatively induced strand breaks in mtDNA has been less thoroughly examined. Polynucleotide kinase/phosphatase (PNKP) processes strand-break termini to render them chemically compatible for the subsequent action of DNA polymerases and ligases. Here, we demonstrate that functionally active full-length PNKP is present in mitochondria as well as nuclei. Downregulation of PNKP results in an accumulation of strand breaks in mtDNA of hydrogen peroxide-treated cells. Full restoration of repair of the H(2)O(2)-induced strand breaks in mitochondria requires both the kinase and phosphatase activities of PNKP. We also demonstrate that PNKP contains a mitochondrial-targeting signal close to the C-terminus of the protein. We further show that PNKP associates with the mitochondrial protein mitofilin. Interaction with mitofilin may serve to translocate PNKP into mitochondria.
Figures

References
-
- Henner WD, Grunberg SM, Haseltine WA. Sites and structure of gamma radiation-induced DNA strand breaks. J. Biol. Chem. 1982;257:11750–11754. - PubMed
-
- Lennartz M, Coquerelle T, Bopp A, Hagen U. Oxygen—effect on strand breaks and specific end-groups in DNA of irradiated thymocytes. Int. J. Radiat. Biol. Relat. Stud. Phys. Chem. Med. 1975;27:577–587. - PubMed
-
- Pourquier P, Pilon AA, Kohlhagen G, Mazumder A, Sharma A, Pommier Y. Trapping of mammalian topoisomerase I and recombinations induced by damaged DNA containing nicks or gaps. Importance of DNA end phosphorylation and camptothecin effects. J. Biol. Chem. 1997;272:26441–26447. - PubMed
-
- Plo I, Liao ZY, Barcelo JM, Kohlhagen G, Caldecott KW, Weinfeld M, Pommier Y. Association of XRCC1 and tyrosyl DNA phosphodiesterase (Tdp1) for the repair of topoisomerase I-mediated DNA lesions. DNA Repair. 2003;2:1087–1100. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

