Identification of RNA-protein interaction networks using PAR-CLIP
- PMID: 22213601
- PMCID: PMC3711140
- DOI: 10.1002/wrna.1103
Identification of RNA-protein interaction networks using PAR-CLIP
Abstract
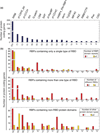
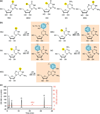
All mRNA molecules are subject to some degree of post-transcriptional gene regulation (PTGR) involving sequence-dependent modulation of splicing, cleavage and polyadenylation, editing, transport, stability, and translation. The recent introduction of deep-sequencing technologies enabled the development of new methods for broadly mapping interaction sites between RNA-binding proteins (RBPs) and their RNA target sites. In this article, we review crosslinking and immunoprecipitation (CLIP) methods adapted for large-scale identification of target RNA-binding sites and the respective RNA recognition elements. CLIP methods have the potential to detect hundreds of thousands of binding sites in single experiments although the separation of signal from noise can be challenging. As a consequence, each CLIP method has developed different strategies to distinguish true targets from background. We focus on photoactivatable ribonucleoside-enhanced CLIP, which relies on the intracellular incorporation of photoactivatable ribonucleoside analogs into nascent transcripts, and yields characteristic sequence changes upon crosslinking that facilitate the separation of signal from noise. The precise knowledge of the position and distribution of binding sites across mature and primary mRNA transcripts allows critical insights into cellular localization and regulatory function of the examined RBP. When coupled with other systems-wide approaches measuring transcript and protein abundance, the generation of high-resolution RBP-binding site maps across the transcriptome will broaden our understanding of PTGR and thereby lead to new strategies for therapeutic treatment of genetic diseases perturbing these processes.
Copyright © 2011 John Wiley & Sons, Ltd.
Figures



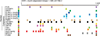
References
-
- Mackay JP, Font J, Segal DJ. The prospects for designer single-stranded RNA-binding proteins. Nat Struct Mol Biol. 2011;18:256–261. - PubMed
-
- Sonnhammer EL, Eddy SR, Durbin R. Pfam: a comprehensive database of protein domain families based on seed alignments. Proteins. 1997;28:405–420. - PubMed
-
- Timchenko LT, Timchenko NA, Caskey CT, Roberts R. Novel proteins with binding specificity for DNA CTG repeats and RNA Cug repeats: implications for myotonic dystrophy. Hum Mol Genet. 1996;5:115–121. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

