S-Nitrosylated proteins in pea (Pisum sativum L.) leaf peroxisomes: changes under abiotic stress
- PMID: 22213812
- PMCID: PMC3295397
- DOI: 10.1093/jxb/err414
S-Nitrosylated proteins in pea (Pisum sativum L.) leaf peroxisomes: changes under abiotic stress
Abstract
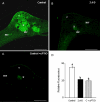
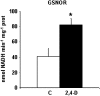
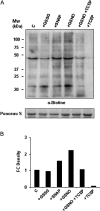
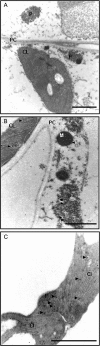
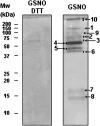
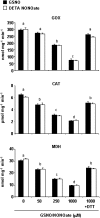
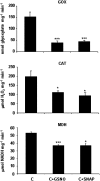
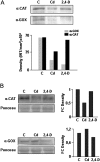
Peroxisomes, single-membrane-bounded organelles with essentially oxidative metabolism, are key in plant responses to abiotic and biotic stresses. Recently, the presence of nitric oxide (NO) described in peroxisomes opened the possibility of new cellular functions, as NO regulates diverse biological processes by directly modifying proteins. However, this mechanism has not yet been analysed in peroxisomes. This study assessed the presence of S-nitrosylation in pea-leaf peroxisomes, purified S-nitrosylated peroxisome proteins by immunoprecipitation, and identified the purified proteins by two different mass-spectrometry techniques (matrix-assisted laser desorption/ionization tandem time-of-flight and two-dimensional nano-liquid chromatography coupled to ion-trap tandem mass spectrometry). Six peroxisomal proteins were identified as putative targets of S-nitrosylation involved in photorespiration, β-oxidation, and reactive oxygen species detoxification. The activity of three of these proteins (catalase, glycolate oxidase, and malate dehydrogenase) is inhibited by NO donors. NO metabolism/S-nitrosylation and peroxisomes were analysed under two different types of abiotic stress, i.e. cadmium and 2,4-dichlorophenoxy acetic acid (2,4-D). Both types of stress reduced NO production in pea plants, and an increase in S-nitrosylation was observed in pea extracts under 2,4-D treatment while no total changes were observed in peroxisomes. However, the S-nitrosylation levels of catalase and glycolate oxidase changed under cadmium and 2,4-D treatments, suggesting that this post-translational modification could be involved in the regulation of H(2)O(2) level under abiotic stress.
Figures



References
-
- Abat JK, Deswal R. Differential modulation of S-nitrosoproteome of Brassica juncea by low temperature: change in S-nitrosylation of Rubisco is responsible for the inactivation of its carboxylase activity. Proteomics. 2009;9:4368–4380. - PubMed
-
- Abat JK, Mattoo AK, Deswal R. S-Nitrosylated proteins of a medicinal CAM plant Kalanchoe pinnata – ribulose-1,5-bisphosphate carboxylase/oxygenase activity targeted for inhibition. FEBS Journal. 2008;275:2862–2872. - PubMed
-
- Aebi H. Catalase in vitro. Methods in Enzymology. 1984;105:121–126. - PubMed
-
- Baker A, Graham I. Plant Peroxisomes. Biochemistry, Cell Biology and Biotechnological Applications. Dordrecht, The Netherlands: Kluwer Academic Publishers; 2002.
-
- Barroso C, Romero LC, Cejudo FJ, Vega JM, Gotor C. Salt-specific regulation of the cytosolic O-acetylserine(thiol) lyase gene from Arabidopsis thaliana is dependent on abscisic acid. Plant Molecular Biology. 1999;40:729–736. - PubMed

