New approaches for dissecting protease functions to improve probe development and drug discovery
- PMID: 22218294
- PMCID: PMC3513415
- DOI: 10.1038/nsmb.2203
New approaches for dissecting protease functions to improve probe development and drug discovery
Abstract
Proteases are well-established targets for pharmaceutical development because of their known enzymatic mechanism and their regulatory roles in many pathologies. However, many potent clinical lead compounds have been unsuccessful either because of a lack of specificity or because of our limited understanding of the biological roles of the targeted protease. In order to successfully develop protease inhibitors as drugs, it is necessary to understand protease functions and to expand the platform of inhibitor development beyond active site-directed design and in vitro optimization. Several newly developed technologies will enhance assessment of drug selectivity in living cells and animal models, allowing researchers to focus on compounds with high specificity and minimal side effects in vivo. In this review, we highlight advances in the development of chemical probes, proteomic methods and screening tools that we feel will help facilitate this paradigm shift in drug discovery.
Figures
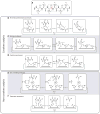

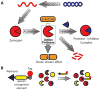

References
-
- Powers JC, Asgian JL, Ekici OD, James KE. Irreversible inhibitors of serine, cysteine, and threonine proteases. Chem Rev. 2002;102:4639–4750. - PubMed
-
- Johnson SL, Pellecchia M. Structure- and fragment-based approaches to protease inhibition. Curr Top Med Chem. 2006;6:317–329. - PubMed
-
- Turk B. Targeting proteases: successes, failures and future prospects. Nat Rev Drug Discov. 2006;5:785–799. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

