Current chemical biology approaches to interrogate protein methyltransferases
- PMID: 22220966
- PMCID: PMC3306480
- DOI: 10.1021/cb200519y
Current chemical biology approaches to interrogate protein methyltransferases
Abstract
Protein methyltransferases (PMTs) play various physiological and pathological roles through methylating histone and nonhistone targets. However, most PMTs including more than 60 human PMTs remain to be fully characterized. The current approaches to elucidate the functions of PMTs have been diversified by many emerging chemical biology technologies. This review focuses on progress in these aspects and is organized into four discussion modules (assays, substrates, cofactors, and inhibitors) that are important to elucidate biological functions of PMTs. These modules are expected to provide general guidance and present emerging methods for researchers to select and combine suitable PMT-activity assays, well-defined substrates, novel SAM surrogates, and PMT inhibitors to interrogate PMTs.
Figures
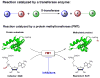




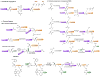


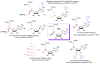
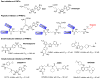
Similar articles
-
Selective inhibitors of protein methyltransferases.J Med Chem. 2015 Feb 26;58(4):1596-629. doi: 10.1021/jm501234a. Epub 2014 Dec 2. J Med Chem. 2015. PMID: 25406853 Free PMC article.
-
Inhibitors of protein methyltransferases as chemical tools.Epigenomics. 2015;7(8):1327-38. doi: 10.2217/epi.15.87. Epub 2015 Dec 8. Epigenomics. 2015. PMID: 26646500 Free PMC article. Review.
-
Formulating a fluorogenic assay to evaluate S-adenosyl-L-methionine analogues as protein methyltransferase cofactors.Mol Biosyst. 2011 Nov;7(11):2970-81. doi: 10.1039/c1mb05230f. Epub 2011 Aug 24. Mol Biosyst. 2011. PMID: 21866297 Free PMC article.
-
Defining efficient enzyme-cofactor pairs for bioorthogonal profiling of protein methylation.Proc Natl Acad Sci U S A. 2013 Oct 15;110(42):16778-83. doi: 10.1073/pnas.1216365110. Epub 2013 Sep 30. Proc Natl Acad Sci U S A. 2013. PMID: 24082136 Free PMC article.
-
Therapeutic targeting potential of the protein lysine and arginine methyltransferases to reverse cancer chemoresistance.Front Mol Biosci. 2024 Dec 5;11:1455415. doi: 10.3389/fmolb.2024.1455415. eCollection 2024. Front Mol Biosci. 2024. PMID: 39703687 Free PMC article. Review.
Cited by
-
Small Molecule Inhibitors of Protein Arginine Methyltransferases.Expert Opin Investig Drugs. 2016;25(3):335-58. doi: 10.1517/13543784.2016.1144747. Epub 2016 Feb 16. Expert Opin Investig Drugs. 2016. PMID: 26789238 Free PMC article. Review.
-
Profiling and Validation of Live-Cell Protein Methylation with Engineered Enzymes and Methionine Analogues.Curr Protoc. 2021 Aug;1(8):e213. doi: 10.1002/cpz1.213. Curr Protoc. 2021. PMID: 34370893 Free PMC article.
-
Protein Arginine Methylation of the Translation Initiation Factor eIF1A Increases Usage of a Near-cognate Start Codon.bioRxiv [Preprint]. 2024 Aug 17:2024.08.16.608280. doi: 10.1101/2024.08.16.608280. bioRxiv. 2024. PMID: 39185183 Free PMC article. Preprint.
-
Bioorthogonal profiling of protein methylation (BPPM) using an azido analog of S-adenosyl-L-methionine.Curr Protoc Chem Biol. 2013;5(1):45-66. doi: 10.1002/9780470559277.ch120240. Curr Protoc Chem Biol. 2013. PMID: 23667794 Free PMC article.
-
SMYD5 catalyzes histone H3 lysine 36 trimethylation at promoters.Nat Commun. 2022 Jun 9;13(1):3190. doi: 10.1038/s41467-022-30940-1. Nat Commun. 2022. PMID: 35680905 Free PMC article.
References
-
- Richon VM, Johnston D, Sneeringer CJ, Jin L, Majer CR, Elliston K, Jerva LF, Scott MP, Copeland RA. Chemogenetic analysis of human protein methyltransferases. Chem Biol Drug Des. 2011;78:199–210. - PubMed
-
- Kouzarides T. SnapShot: Histone-modifying enzymes. Cell. 2007;128:802–803. - PubMed
-
- Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

