Selective analysis of cell-free DNA in maternal blood for evaluation of fetal trisomy
- PMID: 22223233
- PMCID: PMC3500507
- DOI: 10.1002/pd.2922
Selective analysis of cell-free DNA in maternal blood for evaluation of fetal trisomy
Abstract
Objective: To develop a novel prenatal assay based on selective analysis of cell-free DNA in maternal blood for evaluation of fetal Trisomy 21 (T21) and Trisomy 18 (T18).
Methods: Two hundred ninety-eight pregnancies, including 39 T21 and seven T18 confirmed fetal aneuploidies, were analyzed using a novel, highly multiplexed assay, termed digital analysis of selected regions (DANSR™). Cell-free DNA from maternal blood samples was analyzed using DANSR assays for loci on chromosomes 21 and 18. Products from 96 separate patients were pooled and sequenced together. A standard Z-test of chromosomal proportions was used to distinguish aneuploid samples from average-risk pregnancy samples. DANSR aneuploidy discrimination was evaluated at various sequence depths.
Results: At the lowest sequencing depth, corresponding to 204,000 sequencing counts per sample, average-risk cases where distinguished from T21 and T18 cases, with Z statistics for all cases exceeding 3.6. Increasing the sequencing depth to 410,000 counts per sample substantially improved separation of aneuploid and average-risk cases. A further increase to 620,000 counts per sample resulted in only marginal improvement. This depth of sequencing represents less than 5% of that required by massively parallel shotgun sequencing approaches.
Conclusion: Digital analysis of selected regions enables highly accurate, cost efficient, and scalable noninvasive fetal aneuploidy assessment.
© 2012 John Wiley & Sons, Ltd.
Figures



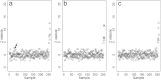
Comment in
-
Noninvasive fetal trisomy 21 detection using chromosome-selective sequencing: a variation of the molecular counting theme.Expert Rev Mol Diagn. 2012 May;12(4):329-31. doi: 10.1586/erm.12.24. Expert Rev Mol Diagn. 2012. PMID: 22616698
References
-
- ACOG Practice Bulletin No. 77. Screening for chromosomal aneuploidies. Obstet Gynecol. 2007;109:217–27. - PubMed
-
- Nicolaides KH. Screening for fetal aneuploidies at 11 to 13 weeks. Prenat Diagn. 2011;31(1):7–15. - PubMed
-
- Caughey AB, Hopkins LM, Norton ME. Chorionic villus sampling compared with amniocentesis and the difference in the rate of pregnancy loss. Obstet Gynecol. 2006;108(3 Pt 1):612–6. - PubMed
-
- Bianchi DW, Simpson JL, Jackson LG, et al. Fetal gender and aneuploidy detection using fetal cells in maternal blood: analysis of NIFTY I data. National Institute of Child Health and Development Fetal Cell Isolation Study. Prenat Diagn. 2002;22(7):609–15. - PubMed
-
- Guetta E, Simchen MJ, Mammon-Daviko K, et al. Analysis of fetal blood cells in the maternal circulation: challenges, ongoing efforts, and potential solutions. Stem Cells Dev. 2004;13(1):93–9. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

