Sialic acid, periodontal pathogens and Tannerella forsythia: stick around and enjoy the feast!
- PMID: 22230462
- PMCID: PMC4049603
- DOI: 10.1111/j.2041-1014.2011.00630.x
Sialic acid, periodontal pathogens and Tannerella forsythia: stick around and enjoy the feast!
Abstract
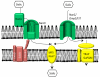
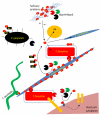
Periodontal pathogens, like any other human commensal or pathogenic bacterium, must possess both the ability to acquire the necessary growth factors and the means to adhere to surfaces or reside and survive in their environmental niche. Recent evidence has suggested that sialic acid containing host molecules may provide both of these requirements in vivo for several periodontal pathogens but most notably for the red complex organism Tannerella forsythia. Several other periodontal pathogens also possess sialic acid scavenging enzymes - sialidases, which can also expose adhesive epitopes, but might also act as adhesins in their own right. In addition, recent experimental work coupled with the release of several genome sequences has revealed that periodontal bacteria have a range of sialic acid uptake and utilization systems while others may also use sialic acid as a cloaking device on their surface to mimic host and avoid immune recognition. This review will focus on these systems in a range of periodontal bacteria with a focus on Ta. forsythia.
© 2011 John Wiley & Sons A/S.
Figures

Similar articles
-
A novel sialic acid utilization and uptake system in the periodontal pathogen Tannerella forsythia.J Bacteriol. 2010 May;192(9):2285-93. doi: 10.1128/JB.00079-10. Epub 2010 Feb 26. J Bacteriol. 2010. PMID: 20190043 Free PMC article.
-
Sialic acid transporter NanT participates in Tannerella forsythia biofilm formation and survival on epithelial cells.Microb Pathog. 2016 May;94:12-20. doi: 10.1016/j.micpath.2015.08.012. Epub 2015 Aug 28. Microb Pathog. 2016. PMID: 26318875 Free PMC article.
-
Evidence for a carbohydrate-binding module (CBM) of Tannerella forsythia NanH sialidase, key to interactions at the host-pathogen interface.Biochem J. 2018 Mar 26;475(6):1159-1176. doi: 10.1042/BCJ20170592. Biochem J. 2018. PMID: 29483296
-
Peptidoglycan synthesis in Tannerella forsythia: Scavenging is the modus operandi.Mol Oral Microbiol. 2018 Apr;33(2):125-132. doi: 10.1111/omi.12210. Epub 2018 Feb 12. Mol Oral Microbiol. 2018. PMID: 29247483 Free PMC article. Review.
-
How bacteria utilize sialic acid during interactions with the host: snip, snatch, dispatch, match and attach.Microbiology (Reading). 2022 Mar;168(3):001157. doi: 10.1099/mic.0.001157. Microbiology (Reading). 2022. PMID: 35316172 Free PMC article. Review.
Cited by
-
Characterization of Porphyromonas gingivalis sialidase and disruption of its role in host-pathogen interactions.Microbiology (Reading). 2019 Nov;165(11):1181-1197. doi: 10.1099/mic.0.000851. Microbiology (Reading). 2019. PMID: 31517596 Free PMC article.
-
Structure of the LPS O-chain from Fusobacterium nucleatum strain 10953, containing sialic acid.Carbohydr Res. 2017 Feb 22;440-441:38-42. doi: 10.1016/j.carres.2017.01.009. Epub 2017 Jan 28. Carbohydr Res. 2017. PMID: 28199859 Free PMC article.
-
Gingipain-dependent degradation of mammalian target of rapamycin pathway proteins by the periodontal pathogen Porphyromonas gingivalis during invasion.Mol Oral Microbiol. 2013 Oct;28(5):366-78. doi: 10.1111/omi.12030. Epub 2013 May 29. Mol Oral Microbiol. 2013. PMID: 23714361 Free PMC article.
-
Glycobiology Aspects of the Periodontal Pathogen Tannerella forsythia.Biomolecules. 2012 Oct 12;2(4):467-82. doi: 10.3390/biom2040467. Biomolecules. 2012. PMID: 24970146 Free PMC article.
-
The use of sialidase therapy for respiratory viral infections.Antiviral Res. 2013 Jun;98(3):401-9. doi: 10.1016/j.antiviral.2013.04.012. Epub 2013 Apr 17. Antiviral Res. 2013. PMID: 23602850 Free PMC article. Review.
References
-
- Agnani G, Tricot-Doleux S, Du L, Bonnaure-Mallet M. Adherence of Porphyromonas gingivalis to gingival epithelial cells: modulation of bacterial protein expression. Oral Microbiol Immunol. 2000;15:48–52. - PubMed
-
- Amith SR, Jayanth P, Franchuk S, et al. Dependence of pathogen molecule-induced toll-like receptor activation and cell function on Neu1 sialidase. Glycoconj J. 2009;26:1197–1212. - PubMed
-
- Amith SR, Jayanth P, Franchuk S, et al. Neu1 desialylation of sialyl alpha-2,3-linked beta-galactosyl residues of TOLL-like receptor 4 is essential for receptor activation and cellular signaling. Cell Signal. 2010;22:314–324. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

