The novel sigma factor-like regulator RpoQ controls luminescence, chitinase activity, and motility in Vibrio fischeri
- PMID: 22233679
- PMCID: PMC3252764
- DOI: 10.1128/mBio.00285-11
The novel sigma factor-like regulator RpoQ controls luminescence, chitinase activity, and motility in Vibrio fischeri
Abstract
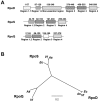
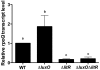
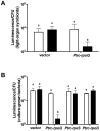
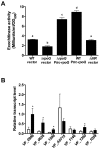
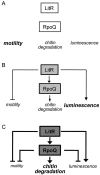
Vibrio fischeri, the bacterial symbiont of the Hawaiian bobtail squid, Euprymna scolopes, uses quorum sensing to control genes involved in bioluminescence, host colonization, and other biological processes. Previous work has shown that AinS/R-directed quorum sensing also regulates the expression of rpoQ (VF_A1015), a gene annotated as an RpoS-like sigma factor. In this study, we demonstrate using phylogenetics that RpoQ is related to, but distinct from, the stationary-phase sigma factor RpoS. Overexpression of rpoQ results in elevated chitinase activity but decreased motility and luminescence, three activities associated with symbiosis. The reduction in bacterial luminescence associated with the overexpression of rpoQ occurs both in culture and within the light-emitting organ of the squid host. This suppression of bioluminescence is due to the repression of the luxICDABEG promoter. Our results highlight RpoQ as a novel regulatory component, embedded in the quorum-signaling network that controls several biological processes in V. fischeri.
Importance: Quorum signaling is a widely occurring phenomenon that functions in diverse bacterial taxa. It is most often found associated with species that interact with animal or plant hosts, either as mutualists or pathogens, and controls the expression of genes critical to tissue colonization. We present the discovery of rpoQ, which encodes a new regulatory component in the quorum-signaling pathway of Vibrio fischeri. RpoQ is a novel protein in the RpoS family of stationary-phase sigma factors. Unlike many other regulatory proteins involved in the quorum-signaling pathways of the Vibrionaceae, the distribution of RpoQ appears to be restricted to only two closely related species. The role of this regulator is to enhance some quorum-signaling outputs (motility) while suppressing others (luminescence). We propose that RpoQ may be a recently evolved or acquired component in V. fischeri that provides this organism with an additional level of regulation to modulate its existing quorum-signaling pathway.
Figures



References
-
- Milton DL. 2006. Quorum sensing in Vibrios: complexity for diversification. Int. J. Med. Microbiol. 296:61–71 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
