Outer membrane protein A and OprF: versatile roles in Gram-negative bacterial infections
- PMID: 22240162
- PMCID: PMC3338869
- DOI: 10.1111/j.1742-4658.2012.08482.x
Outer membrane protein A and OprF: versatile roles in Gram-negative bacterial infections
Abstract
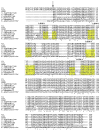
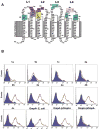
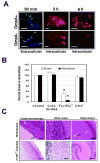
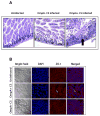
Outer membrane protein A (OmpA) is an abundant protein of Escherichia coli and other enterobacteria and has a multitude of functions. Although the structural features and porin function of OmpA have been well studied, its role in the pathogenesis of various bacterial infections has emerged only during the last decade. The four extracellular loops of OmpA interact with a variety of host tissues for adhesion to and invasion of the cell and for evasion of host-defense mechanisms when inside the cell. This review describes how various regions present in the extracellular loops of OmpA contribute to the pathogenesis of neonatal meningitis induced by E. coli K1 and to many other functions. In addition, the function of OmpA-like proteins, such as OprF of Pseudomonas aeruginosa, is discussed.
© 2012 The Authors Journal compilation © 2012 FEBS.
Figures

Similar articles
-
Escherichia coli outer membrane protein A adheres to human brain microvascular endothelial cells.Biochem Biophys Res Commun. 2005 May 20;330(4):1199-204. doi: 10.1016/j.bbrc.2005.03.097. Biochem Biophys Res Commun. 2005. PMID: 15823570
-
Identification of minimum carbohydrate moiety in N-glycosylation sites of brain endothelial cell glycoprotein 96 for interaction with Escherichia coli K1 outer membrane protein A.Microbes Infect. 2014 Jul;16(7):540-52. doi: 10.1016/j.micinf.2014.06.002. Epub 2014 Jun 13. Microbes Infect. 2014. PMID: 24932957 Free PMC article.
-
Experimental validation of the predicted binding site of Escherichia coli K1 outer membrane protein A to human brain microvascular endothelial cells: identification of critical mutations that prevent E. coli meningitis.J Biol Chem. 2010 Nov 26;285(48):37753-61. doi: 10.1074/jbc.M110.122804. Epub 2010 Sep 17. J Biol Chem. 2010. PMID: 20851887 Free PMC article.
-
The OmpA family of proteins: roles in bacterial pathogenesis and immunity.Vet Microbiol. 2013 May 3;163(3-4):207-22. doi: 10.1016/j.vetmic.2012.08.019. Epub 2012 Aug 31. Vet Microbiol. 2013. PMID: 22986056 Review.
-
OmpA: gating and dynamics via molecular dynamics simulations.Biochim Biophys Acta. 2008 Sep;1778(9):1871-80. doi: 10.1016/j.bbamem.2007.05.024. Epub 2007 Jun 2. Biochim Biophys Acta. 2008. PMID: 17601489 Review.
Cited by
-
High Risk Clone: A Proposal of Criteria Adapted to the One Health Context with Application to Enterotoxigenic Escherichia coli in the Pig Population.Antibiotics (Basel). 2021 Feb 28;10(3):244. doi: 10.3390/antibiotics10030244. Antibiotics (Basel). 2021. PMID: 33671102 Free PMC article.
-
Preliminary Extraction and Identification of the 44.5 kDa Outer Membrane Proteins Isolated from Bovine Fusobacterium necrophorum (AB).Indian J Microbiol. 2013 Dec;53(4):395-9. doi: 10.1007/s12088-013-0388-x. Epub 2013 Mar 16. Indian J Microbiol. 2013. PMID: 24426142 Free PMC article.
-
The Analysis of OmpA and Rz/Rz1 of Lytic Bacteriophage from Surabaya, Indonesia.Scientifica (Cairo). 2021 Dec 23;2021:7494144. doi: 10.1155/2021/7494144. eCollection 2021. Scientifica (Cairo). 2021. PMID: 35096434 Free PMC article.
-
Interaction of an Acinetobacter baumannii's Membrane Protein with Human Fibronectin to Evade Immune Response.Chemistry. 2025 Aug 13;31(45):e00874. doi: 10.1002/chem.202500874. Epub 2025 Jul 25. Chemistry. 2025. PMID: 40708401 Free PMC article.
-
Development of a chitosan-modified PLGA nanoparticle vaccine for protection against Escherichia coli K1 caused meningitis in mice.J Nanobiotechnology. 2021 Mar 5;19(1):69. doi: 10.1186/s12951-021-00812-9. J Nanobiotechnology. 2021. PMID: 33673858 Free PMC article.
References
-
- Koebnik R, Locher KP, Van Gelder P. Structure and function of bacterial outer membrane proteins: barriers in a nutshell. Mol Microbiol. 2000;37:239–253. - PubMed
-
- Smith SG, Mahon V, Lambert MA, Fagan RP. A molecular Swiss army knife: OmpA structure, function and expression. FEMS Microbiol Lett. 2007;273:1–11. - PubMed
-
- Rooijakkers SH, van Strijp JA. Bacterial complement evasion. Mol Immunol. 2007;44:23–32. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

