Functional specialization of the plant miR396 regulatory network through distinct microRNA-target interactions
- PMID: 22242012
- PMCID: PMC3252272
- DOI: 10.1371/journal.pgen.1002419
Functional specialization of the plant miR396 regulatory network through distinct microRNA-target interactions
Abstract
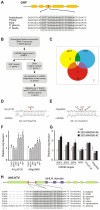
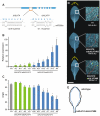
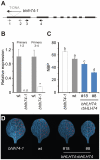
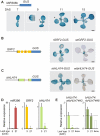
MicroRNAs (miRNAs) are ∼21 nt small RNAs that regulate gene expression in animals and plants. They can be grouped into families comprising different genes encoding similar or identical mature miRNAs. Several miRNA families are deeply conserved in plant lineages and regulate key aspects of plant development, hormone signaling, and stress response. The ancient miRNA miR396 regulates conserved targets belonging to the GROWTH-REGULATING FACTOR (GRF) family of transcription factors, which are known to control cell proliferation in Arabidopsis leaves. In this work, we characterized the regulation of an additional target for miR396, the transcription factor bHLH74, that is necessary for Arabidopsis normal development. bHLH74 homologs with a miR396 target site could only be detected in the sister families Brassicaceae and Cleomaceae. Still, bHLH74 repression by miR396 is required for margin and vein pattern formation of Arabidopsis leaves. MiR396 contributes to the spatio-temporal regulation of GRF and bHLH74 expression during leaf development. Furthermore, a survey of miR396 sequences in different species showed variations in the 5' portion of the miRNA, a region known to be important for miRNA activity. Analysis of different miR396 variants in Arabidopsis thaliana revealed that they have an enhanced activity toward GRF transcription factors. The interaction between the GRF target site and miR396 has a bulge between positions 7 and 8 of the miRNA. Our data indicate that such bulge modulates the strength of the miR396-mediated repression and that this modulation is essential to shape the precise spatio-temporal pattern of GRF2 expression. The results show that ancient miRNAs can regulate conserved targets with varied efficiency in different species, and we further propose that they could acquire new targets whose control might also be biologically relevant.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
miR396a-Mediated basic helix-loop-helix transcription factor bHLH74 repression acts as a regulator for root growth in Arabidopsis seedlings.Plant Cell Physiol. 2014 Jul;55(7):1343-53. doi: 10.1093/pcp/pcu058. Epub 2014 May 3. Plant Cell Physiol. 2014. PMID: 24793750
-
Control of cell proliferation in Arabidopsis thaliana by microRNA miR396.Development. 2010 Jan;137(1):103-12. doi: 10.1242/dev.043067. Development. 2010. PMID: 20023165 Free PMC article.
-
Ectopic expression of miR396 suppresses GRF target gene expression and alters leaf growth in Arabidopsis.Physiol Plant. 2009 Jun;136(2):223-36. doi: 10.1111/j.1399-3054.2009.01229.x. Epub 2009 Feb 2. Physiol Plant. 2009. PMID: 19453503
-
Basic Helix-Loop-Helix (bHLH) Transcription Factors Regulate a Wide Range of Functions in Arabidopsis.Int J Mol Sci. 2021 Jul 1;22(13):7152. doi: 10.3390/ijms22137152. Int J Mol Sci. 2021. PMID: 34281206 Free PMC article. Review.
-
Control of cell proliferation by microRNAs in plants.Curr Opin Plant Biol. 2016 Dec;34:68-76. doi: 10.1016/j.pbi.2016.10.003. Epub 2016 Oct 26. Curr Opin Plant Biol. 2016. PMID: 27794260 Review.
Cited by
-
High-throughput sequencing of small RNA transcriptome reveals salt stress regulated microRNAs in sugarcane.PLoS One. 2013;8(3):e59423. doi: 10.1371/journal.pone.0059423. Epub 2013 Mar 27. PLoS One. 2013. PMID: 23544066 Free PMC article.
-
Potassium Deficiency Significantly Affected Plant Growth and Development as Well as microRNA-Mediated Mechanism in Wheat (Triticum aestivum L.).Front Plant Sci. 2020 Aug 14;11:1219. doi: 10.3389/fpls.2020.01219. eCollection 2020. Front Plant Sci. 2020. PMID: 32922417 Free PMC article.
-
Genome-wide identification and expression analysis of growth-regulating factors in Dendrobium officinale and Dendrobium chrysotoxum.PeerJ. 2023 Dec 15;11:e16644. doi: 10.7717/peerj.16644. eCollection 2023. PeerJ. 2023. PMID: 38111654 Free PMC article.
-
Differential methylation during maize leaf growth targets developmentally regulated genes.Plant Physiol. 2014 Mar;164(3):1350-64. doi: 10.1104/pp.113.233312. Epub 2014 Jan 31. Plant Physiol. 2014. PMID: 24488968 Free PMC article.
-
Divergence in Patterns of Leaf Growth Polarity Is Associated with the Expression Divergence of miR396.Plant Cell. 2015 Oct;27(10):2785-99. doi: 10.1105/tpc.15.00196. Epub 2015 Sep 26. Plant Cell. 2015. PMID: 26410303 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

